Locus Rank
39Gene
Id
183Gene Name
CACNA1IDuplicated
FalseMaternal
TrueGene Not Within Locus (Nearby
FalseGene Description
This gene encodes the pore-forming alpha subunit of a voltage gated calcium channel. The encoded protein is a member of a subfamily of calcium channels referred to as is a low voltage-activated, T-type, calcium channel. The channel encoded by this protein is characterized by a slower activation and inactivation compared to other T-type calcium channels. This protein may be involved in calcium signaling in neurons. Mouse Phenotype
Additional Image
Allen Brain
Allen Regions
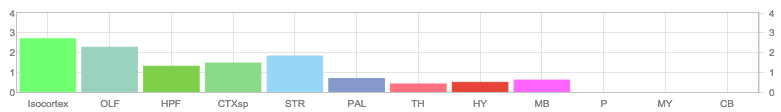
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
http://zfin.org/ZDB-GENE-060503-324Allen Link
http://mouse.brain-map.org/gene/show/88588More Additional Images
Papers
Locus Rank
Locus Rank
39Genes in Region
CACNA1IAssociated Snps
chr22_39987017_DGWAS Region (hg19)
chr22:39975317-40016817Genes Skipping
Ricopili Plot Surrounding Area
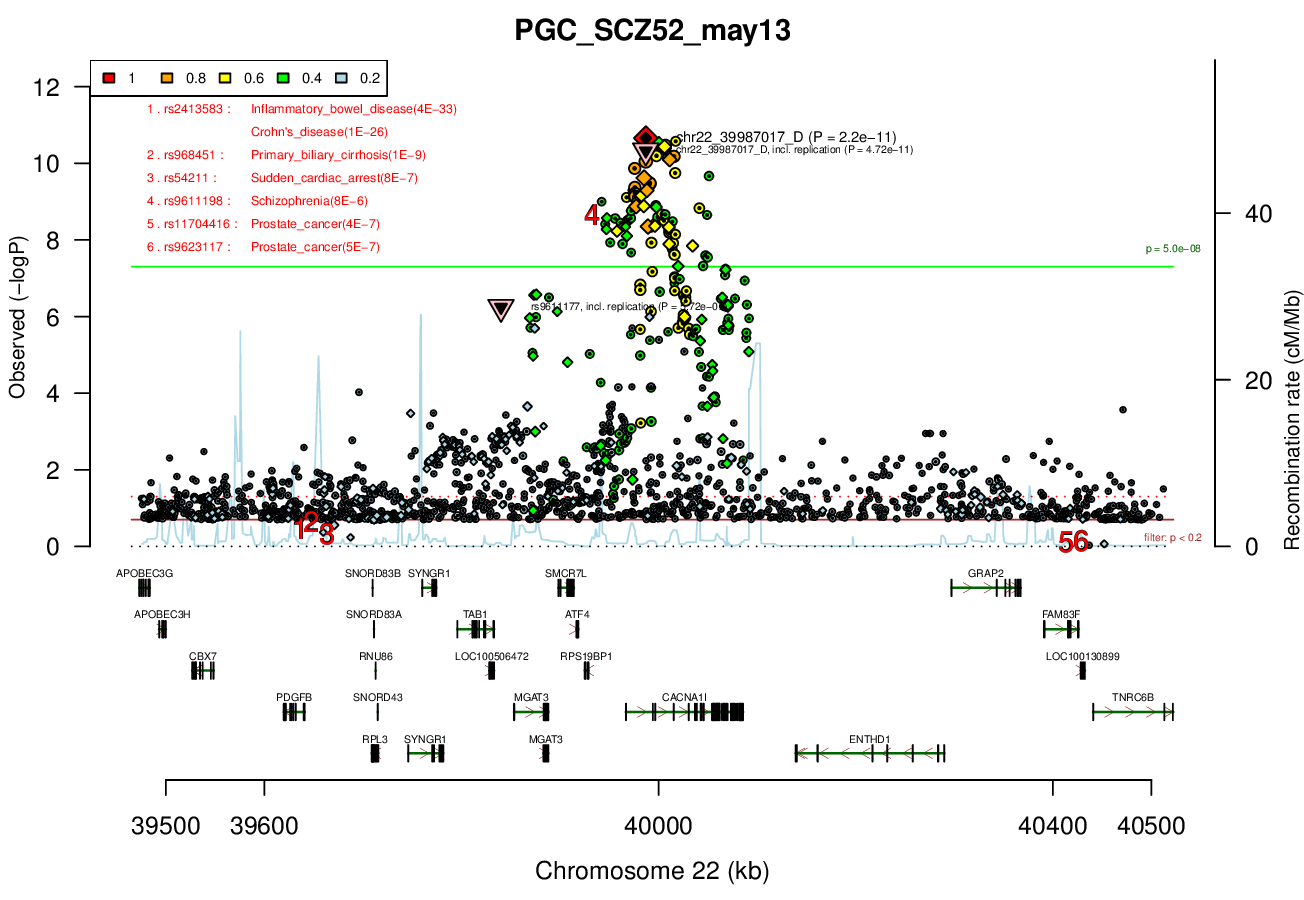
Distance On Each Side Of Locus In Above Plot
0.5 MBRicopili Plot
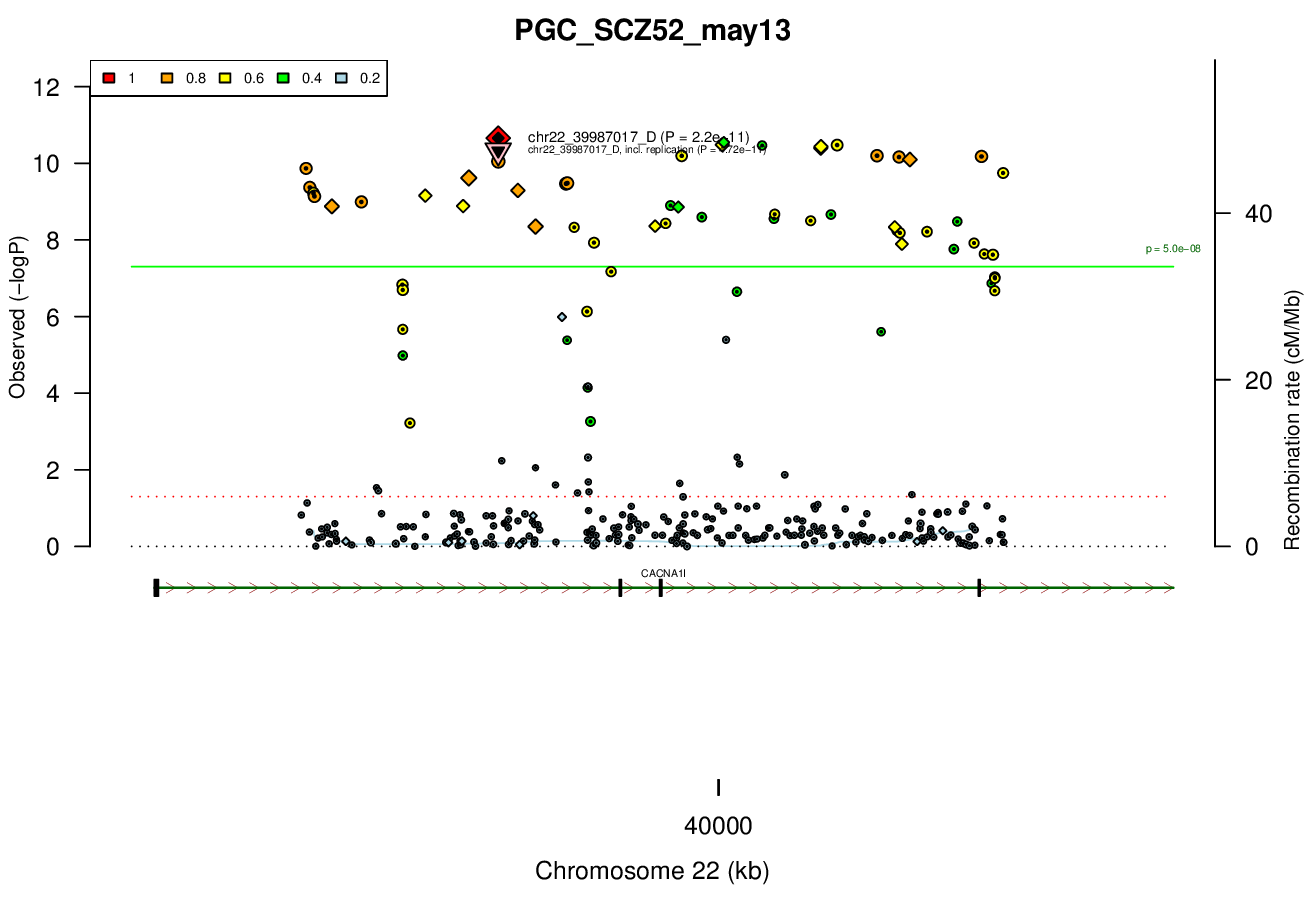
Gwas And Surrounding Region Pic
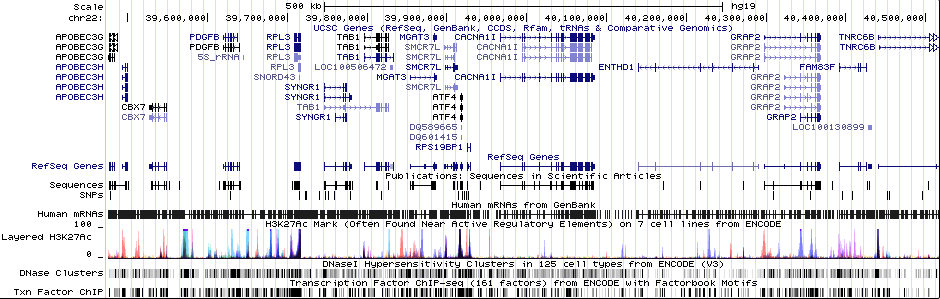
Gwas Region Pic
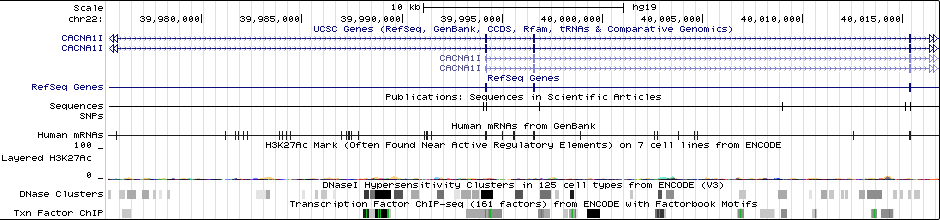
Sequence 1
Ensembl Gene Name
cacna1iHarvard Allele
a245Mutation Area - WT DNA Sequence
CACCCTTCCCACACCATATTATCAGCCTGAGGAGGATGACGAGCGGCCCTTCATCTGCTCGTTGGCTCAAGATAACGGCATCATGTCCTGCAGCGATGTGCCTGCTCGGAGAGAGGGCGGACGGGAGTGCTGTCTGGATAAAGAGGACGCTCTTCACCGGCAGGCTCTGGGTCTGAGCGCCGAGCCGCTGGTCAACGGCAGCGCATCTGCCATGGGCCTCTGTGTGAACTGGAATCAGTACTACACCCGCTGCCACACTGGACACACCAATCCTCATAAAGGAGCAATTAATTTCGACAACATAGGATACGCATGGATCMutation Area - Mutant DNA Sequence
CACCCTTCCCACACCATATTATCAGCCTGAGGAGGATGACGAGCGGCCCTTCATCGTCTGTGTGAACTGGAATCAGTACTACACCCGCTGTCTGGATACACACTGGACACACCAATCCTCATAAAGGAGCAATTAATTTCGACAACATAGGATACGCATGGATCGuide RNA Target Sites
GCGGCCCTTCATCTGCTCGTTGG
TGCTCGGAGAGAGGGCGGACGGG
CCAGTTCACACAGAGGCCCATGG
GGTGTGTCCAGTGTGGCAGCGGG
Genotyping Primers
f, CACCCTTCCCACACCATATTATCAGC
r, GATCCATGCGTATCCTATGTTGTCG
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
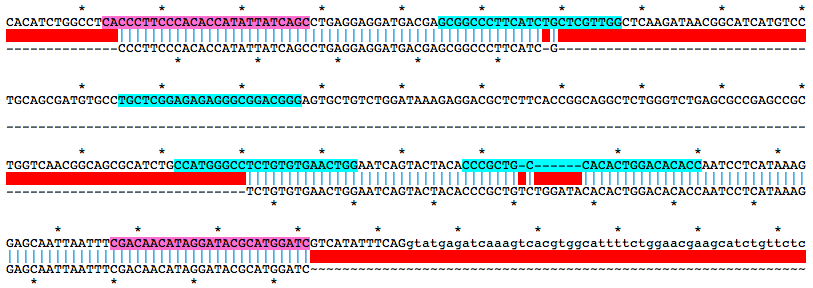
Allele
162bpD,7bpIWT Genotyping Size
319WT Protein Sequence
MTTEECPVPLGVSQVGAGLCEDFGPSQGDDIIEEVGPCDDTGSDLSTLVVPFPELAPVVF
FCLKQTTCPRNWCIRMVWFERISIMVILLNCVTLGMYQPCENIDCTSERCQVLQAFDAFI
YIFFALEMVVKMVALGIFGRRCYLGDTWNRLDFFIVMAGMVEYSLDLQNINFSAIRTVRV
LRPLKAINRVPSMRILVNLLLDTLPMLGNVLLLCFFVFFIFGIIGVQLWAGLLRNRCYPE
ENFTLTSGLTLPTPYYQPEEDDERPFICSLAQDNGIMSCSDVPARREGGRECCLDKEDAL
HRQALGLSAEPLVNGSASAMGLCVNWNQYYTRCHTGHTNPHKGAINFDNIGYAWIVIFQV
ITLEGWVEIMYYVMDAHSFYNFIYFIFLIIIGSFFMINLCLVVIATQFSETKQREHQLMQ
EQRARYLSSSTLASLAEPGDCYEELFQLVCHILRKARRRSAALYYMLRGKAPPPGGGRGR
GKGGAGSIGGGANVNGGEKHHCHSAQTSHCIHETKLDHSSENAVSLSISPNPEECPQCAA
ALSLKEGEESTGHSANGEEEEGALEETDRDENRLDKKRSSDSDDTLKKKRTCLGKCKDVW
DEIRVKLWGIVESKYFNRGIMIAILINTISMGIEHHNQPDELTNVLEICNIVFTSMFTLE
MILKLTAFGFFEYLRNPYNIFDGIIVIISVCEIIGQSDGGLSVLRTFRLLRVIKLVRFMP
ALRRQLVVLMKTMDNVATFCMLLMLFIFIFSILGMHIFGCKFSLKTEAGDTVPDRKNFDS
LLWAIVTVFQILTQEDWNMVLYNGMASTSPLAALYFVALMTFGNYVLFNLLVAILVEGFQ
AEGDANRSYSDDDRSSCNLEETEKKDSLQLSDPKISTLTPNGHLDLAPAPNARGFYPSER
LSFALGSRKSSVSSLGRASLEQTTLCPGRASLYHNWGRPARPGIWSRRSSWNSLGRSSRC
LGMGGGGSLRVRSPHCHPAEQESLLSPPPPPLHPPLLPRHFPPRRERRALSLELPELLQV
PGPPLPPLHPRQRKKSFSGGLGSVGEHQDCNGKTPSVQPQIINEVYPQVNTRKDREDLED
ELDYSLCFRIQKMMEVYRPDWCETREDWSVFLFSPQNKFRLLCQSIIAHKLFDYVVLAFI
FSNCITVALERPKILQGSLERLFLTVSNYIFTAIFVGEMTLKVVSMGLYIGEQAYLRSSW
NILDGFLVFVSLIDIVVSMAGGAKILGVLRVLRLLRTLRPLRVISRAPGLKLVVETLITS
LKPIGNIVLICCAFFIIFGILGVQLFKGKFYYCLGLDVKNITNKSDCLLANYKWVHHKYN
FDNLGQALMSLFVLASKDGWVNIMYHGLDAVAVDQQPITNNNPWMLLYFISFLLIVSFFV
LNMFVGVVVENFHKCRQHQEVEEAKRREEKRQRRMEKKRRKAQKLPYYASYSHVRLMIHT
LCTSHYLDIFITFIICVNVVTMSLEHYSQPHSLEIALKYCNYFFTSTFVLEAVLKLIAFG
FRRFFKDRWNQLDLAIVLLSVMGITLEEIEISAALPINPTIIRIMRVLRIARVLKLLKMA
TGMRALLDTVVQALPQVGNLGLLFMLLFFIYAALGVELFGELVCNEDYPCEGMSRHATFE
NFGMAFLTLFQVSTGDNWNGIMKDTLRECPPGEYTCNPSLQFISPLYFVSFVLTAQFVLI
NVVVAVLMKHLDDSNKEAQEEAEMDAEIEMELAQGTLCCIGGGVAVADRTGMGHQGAASC
SVVPPHSPITHPVAHEPNSIRRLYSPAQENLWLDSVSLLIKDSFEGEMLMIDNLSGSVFH
HYSSPPVCKDCRTHPQEQIHMAELEQASLRSEQLSDKSSSPALPDDLSLDEQSVYQMAAK
EGKERVCDECHHSEEPGGRGNSRLRRSSRVHSSGTEDGGTCQSPHHNTPARHSIGGVPCS
SSSQEHPGVSGGSSRSNTPVCLPAEFFHPAAAALPAPPRGRTGHKPRGLRLTSPASWASL
RSPGANTRLLTTQYPSHSDSSLATGSSEGSLPTTMEEGLSFIVSTPQPLLDTLLDERTSL
STETLRPSPPATHILQTTRGHQRSRSSCETDINTGSIRQGSEADVGLVWCSSEQTCDSQQ
LSETQSSLSLTSLLLPSSLVPLSVKKCNSTGSLDQGTLTAQEKVFTKEPQGNITSPWKER
KERQSEAAQGTDGESTSQITIAGSRKNR-
Mutant Protein Sequence
MTTEECPVPLGVSQVGAGLCEDFGPSQGDDIIEEVGPCDDTGSDLSTLVVPFPELAPVVF
FCLKQTTCPRNWCIRMVWFERISIMVILLNCVTLGMYQPCENIDCTSERCQVLQAFDAFI
YIFFALEMVVKMVALGIFGRRCYLGDTWNRLDFFIVMAGMVEYSLDLQNINFSAIRTVRV
LRPLKAINRVPSMRILVNLLLDTLPMLGNVLLLCFFVFFIFGIIGVQLWAGLLRNRCYPE
ENFTLTSGLTLPTPYYQPEEDDERPFIVCVNWNQYYTRCLDTHWTHQSS-
Mutant Genotyping Size
164Ribosome Profiling Development
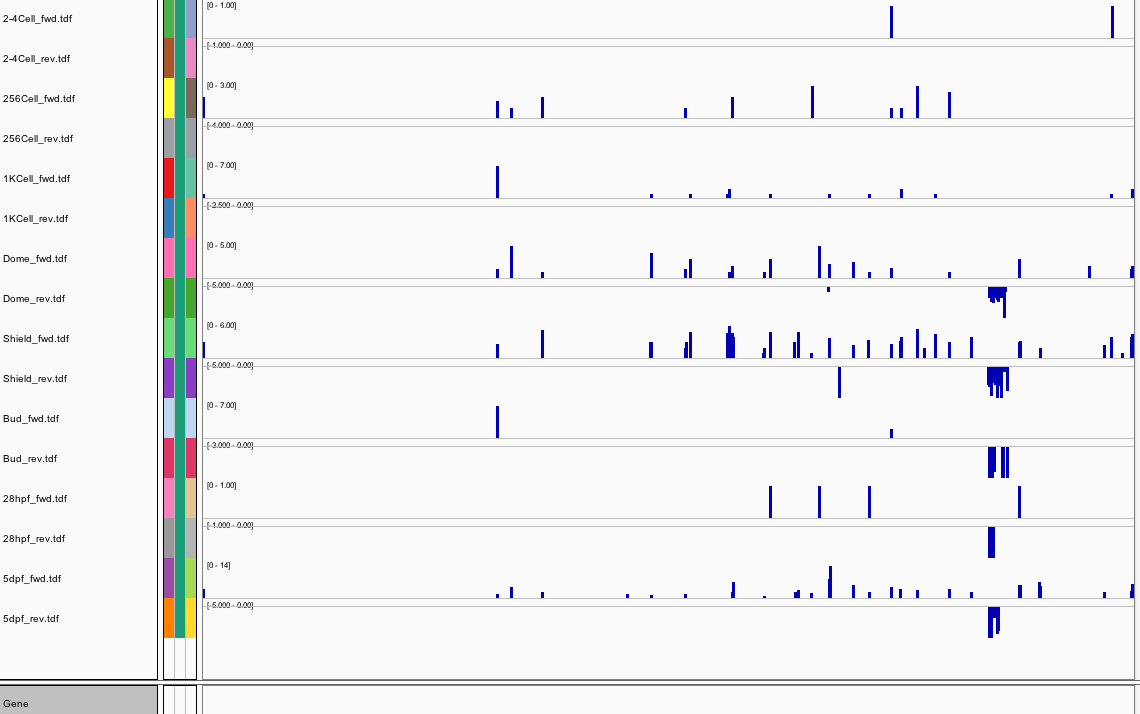
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = not significant 41 het vs 38 hom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
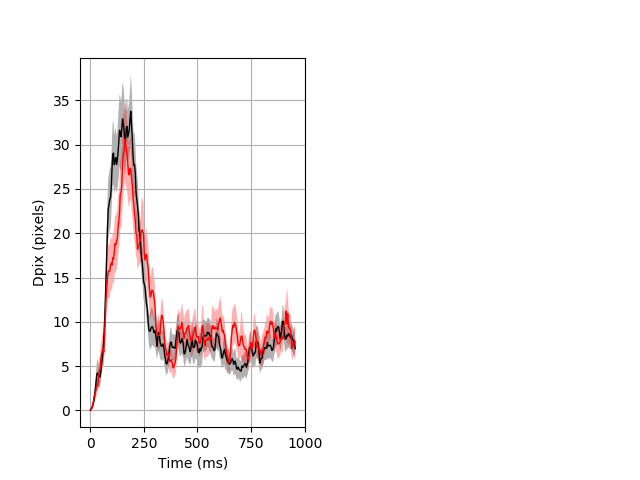
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0381 37 het vs 53 hom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
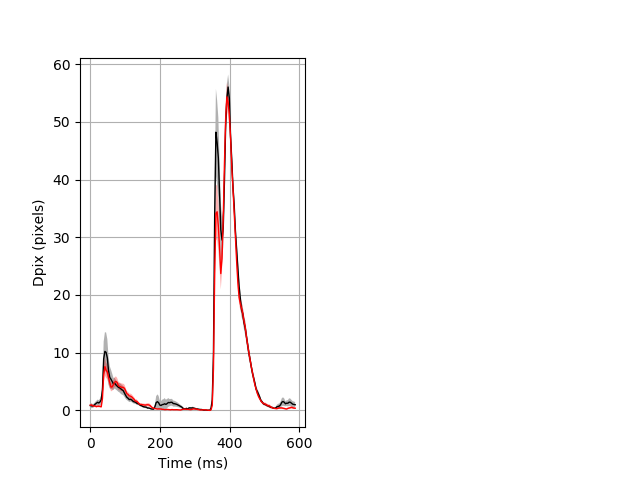
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 31 het vs 22 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
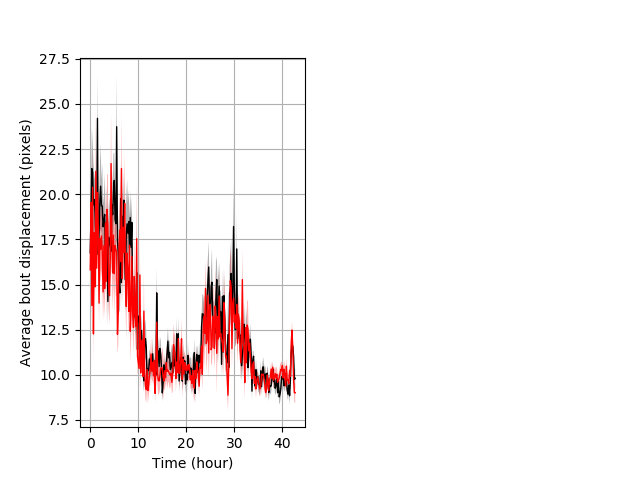
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 41 het vs 38 hom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
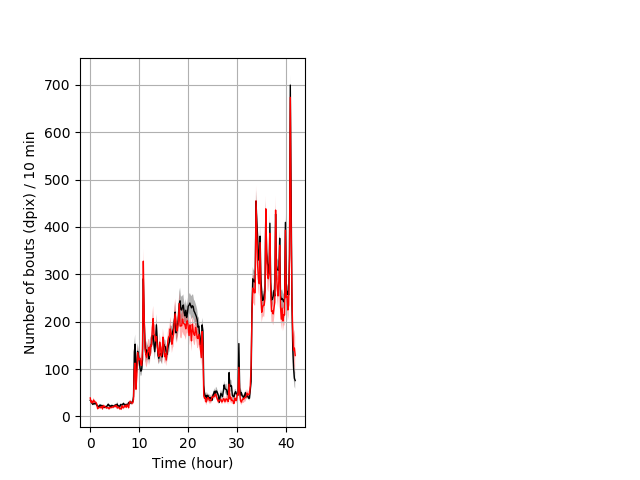
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 37 het vs 53 hom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
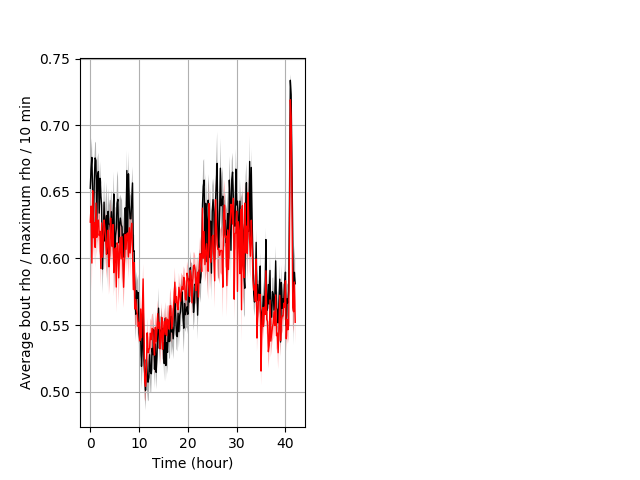
Behavior Data Description 6
Features of movement Graph Pvalue = 0.001 41 het vs 38 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_day2heatshock.pngBehavior Data Graph 6
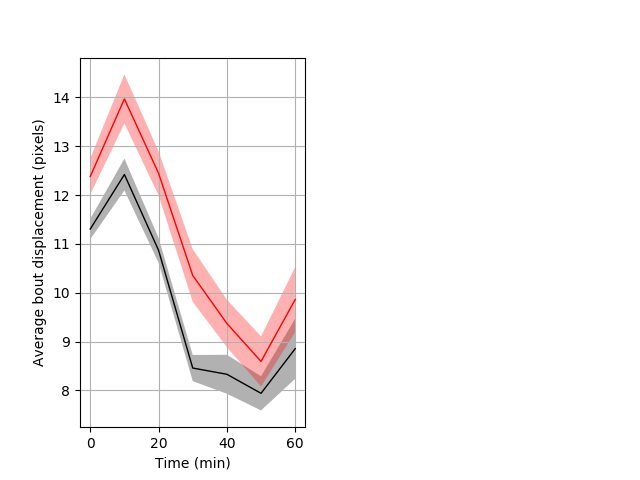
Behavior Data Description 7
PPI tap Graph Pvalue = 0.00156 37 het vs 53 hom boxgraph_pd__smallmovesribgraph_mean_ribbon_totaldistanceresponse_dist_nightprepulseinhibition100b.pngBehavior Data Graph 7
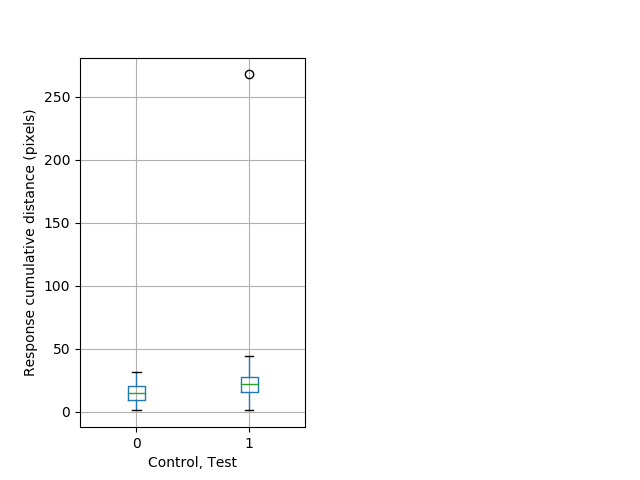
Behavior Data Description 8
Strong tap Graph Pvalue = 0.00952 37 het vs 53 hom boxgraph_pd__ribgraph_mean_ribbon_fullboutdatamax_dpix_adaytappostbdaytappre102.pngBehavior Data Graph 8
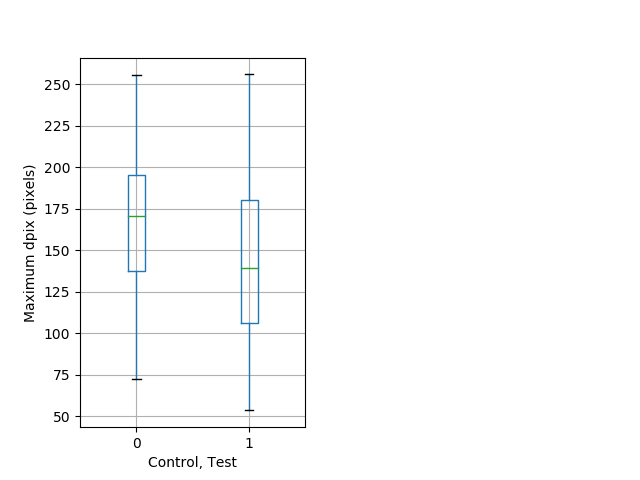
Behavior Data Description 9
Weak tap Graph Pvalue = 0.011 37 het vs 53 hom boxgraph_pd__smallmovesribgraph_mean_ribbon_freqresponse_dpix_shortnightprepulseinhibition100a.pngBehavior Data Graph 9
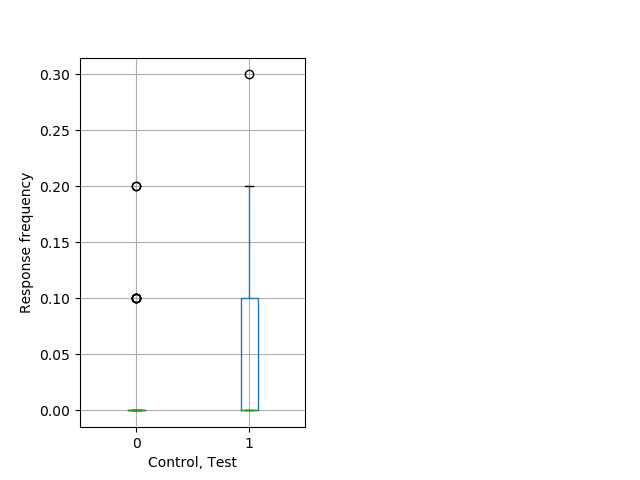
Behavior Data Description 10
NoneBehavior Data Graph 10
NoneBehavior Data Description 11
NoneBehavior Data Graph 11
NoneBehavior Data Description 12
NoneBehavior Data Graph 12
NoneBehavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None