Locus Rank
82Gene
Id
234Gene Name
GRIN2ADuplicated
TrueMaternal
TrueGene Not Within Locus (Nearby
FalseGene Description
This gene encodes a member of the glutamate-gated ion channel protein family. The encoded protein is an N-methyl-D-aspartate (NMDA) receptor subunit. NMDA receptors are both ligand-gated and voltage-dependent, and are involved in long-term potentiation, an activity-dependent increase in the efficiency of synaptic transmission thought to underlie certain kinds of memory and learning. These receptors are permeable to calcium ions, and activation results in a calcium influx into post-synaptic cells, which results in the activation of several signaling cascades. Disruption of this gene is associated with focal epilepsy and speech disorder with or without mental retardation. Mouse Phenotype
Additional Image
Allen Brain
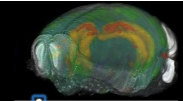
Allen Regions
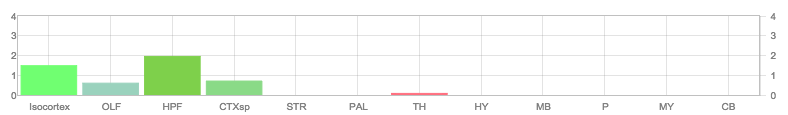
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
http://zfin.org/search?q=grin2a&category=Gene+%2F+Transcript&fq=type%3A%22Gene%22Allen Link
http://mouse.brain-map.org/experiment/show/71307394More Additional Images
Papers
http://www.ncbi.nlm.nih.gov/pubmed/25224260
http://www.ncbi.nlm.nih.gov/pubmed/25159198
Locus Rank
Locus Rank
82Genes in Region
GRIN2AAssociated Snps
rs9922678GWAS Region (hg19)
chr16:9875519-9970219Genes Skipping
Ricopili Plot Surrounding Area
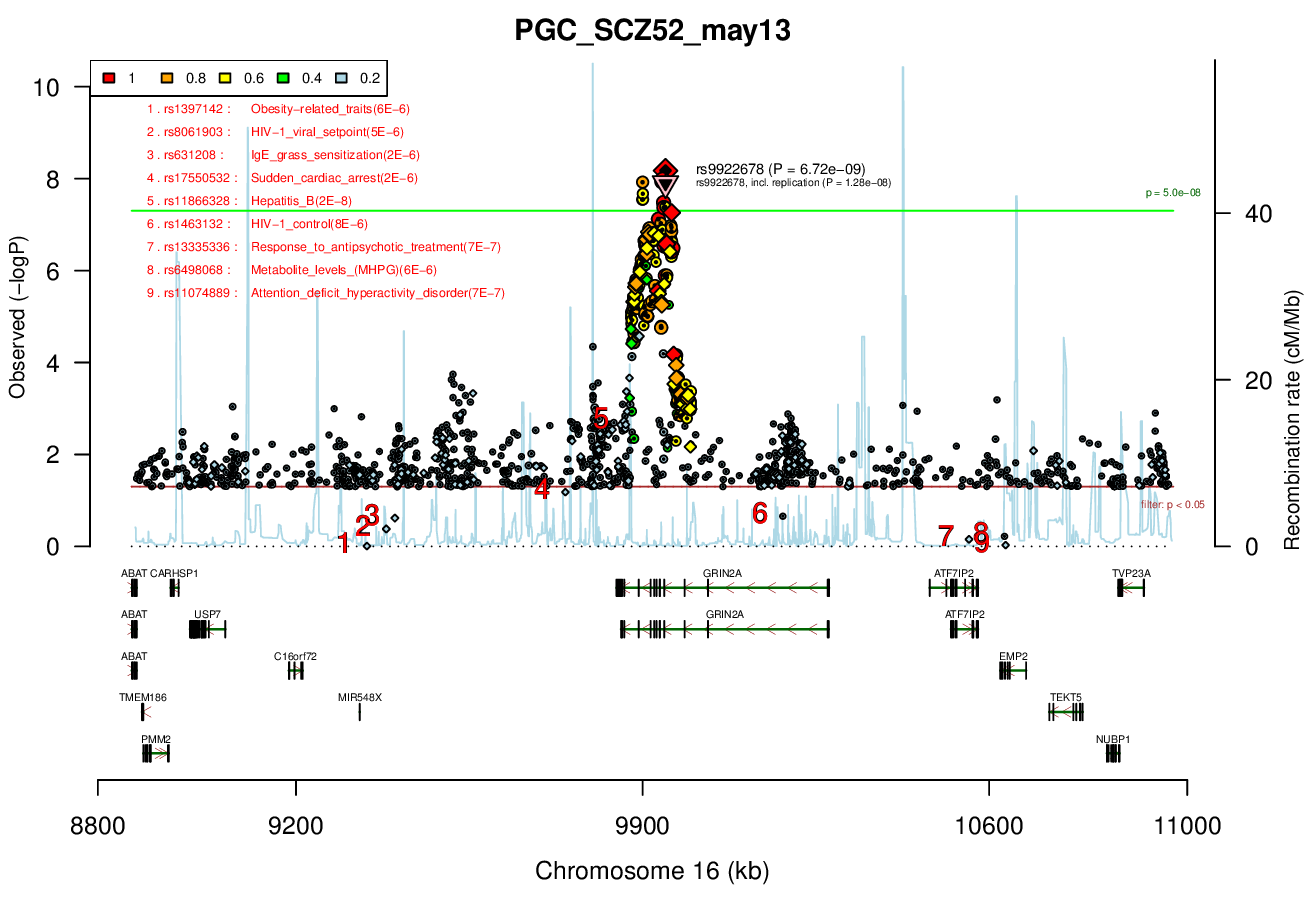
Distance On Each Side Of Locus In Above Plot
1 MBRicopili Plot
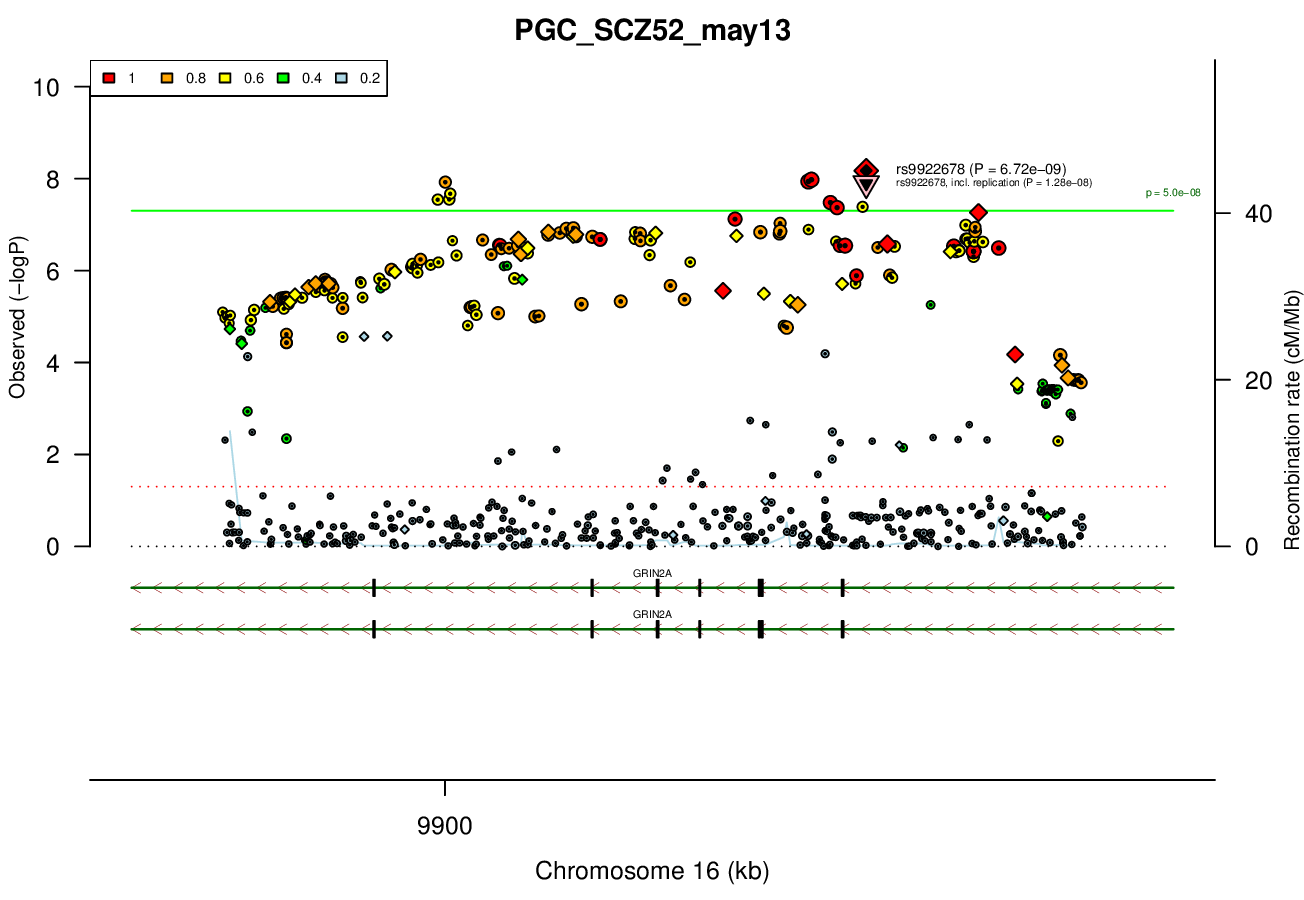
Gwas And Surrounding Region Pic
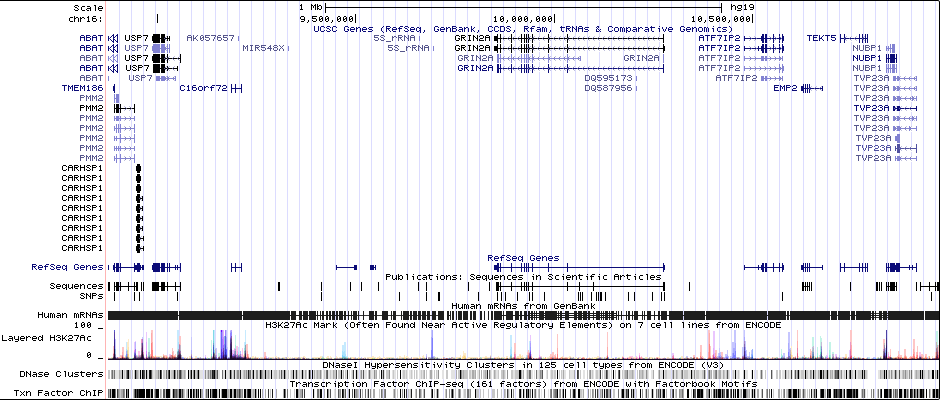
Gwas Region Pic
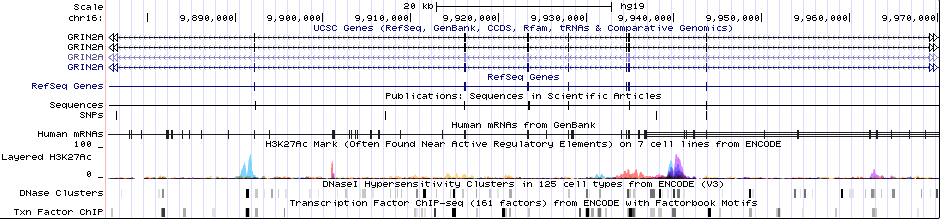
Sequence 1
Ensembl Gene Name
grin2aaHarvard Allele
a280Mutation Area - WT DNA Sequence
GAAGTCGAGCACTATGGGGTTTTCCAGGTTGGTTCTTCTGACTCTTCCAGCTCTCCTTACCCTCATCAGCATCTCACATGCGGCAGAGAAAGTGCCCATACTGAACATTGCTGTGATCCTGGGTCAGACTCGCTACATCTCTGACAGAGATATACGGGCTCTGTGGAGCAAAGATGATCCTATTGATGTCAATGTGGTGACCCTGCTGGTGAATGAAACAGACCCCAAAAGCATCATTACCCATGTGTGTGACCTCATGTCCGGGACTAAAATCCACGGGGTGGTGTTCGGGGACGGCACAGACCAGGAAGCCATTGCCCAGATTTTGGATTTTATTTCCTCTCAGACTTTGATACCCATCCTTGGGATCCATGGTGGCTCATCCATGATCATGGCGGATAAGgtcagtagtatgccaagcatcagctcaactgctcaaaaagatgagcgggaaatattattcacaaaatgagttctgctggtaaaagtatgcgcccttaatgtggtttggtaatgacaatgatMutation Area - Mutant DNA Sequence
GAAGTCGAGCACTATGGGGTTTTCCAGGTTGGTTCTTCTGACTCTTCCAGCTCTCCTTACCCTCATCAGCATCTCACATGCGGCAGAGAAAGTGCCCATACTGAACATTGCTGTGATCCTGGGTCAGACTCGCTACATCTCTGACAGAGATATACGGGCTCTGTGGGGCAAAGATGATCCTATTGATGAACATTGGCACAGACCAGGAAGCCATTGCCCAGATTTTGGATTTTATTTCCTCTCAGACTTTGATACCCATCCTTGGGATCCATGGTGGCTCATCCATGATCATGGCGGATAAGgtcagtagtatgccaagcatcagctcaactgctcaaaaagatgagcgggaaatattattcacaaaatgagttctgctggtaaaagtatgcgcccttaatgtggtttggtaatgacaatgatGuide RNA Target Sites
GGACATGAGGTCACACACATGGGGenotyping Primers
f, GAAGTCGAGCACTATGGGGTTTTCC
r, atcattgtcattaccaaaccacattaaggg
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
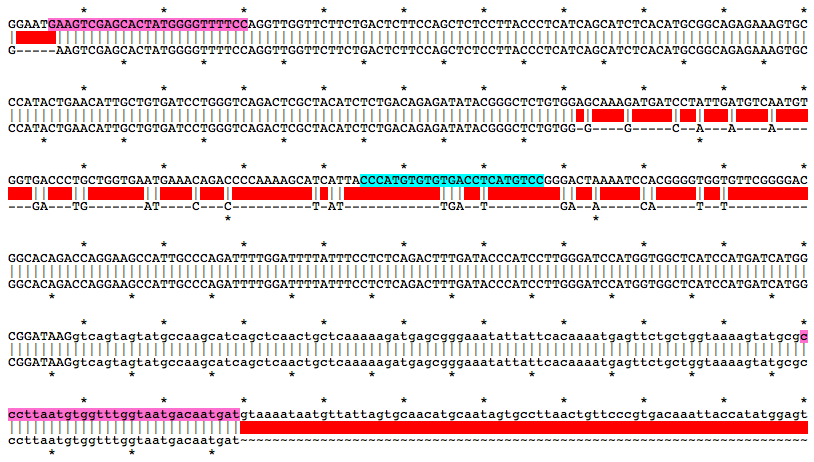
Allele
101bpDWT Genotyping Size
524WT Protein Sequence
MKSSTMGFSRLVLLTLPALLTLISISHAAEKVPILNIAVILGQTRYISDRDIRALWSKDD
PIDVNVVTLLVNETDPKSIITHVCDLMSGTKIHGVVFGDGTDQEAIAQILDFISSQTLIP
ILGIHGGSSMIMADKDVKSTFFQFGASIQQEALLMLNIMEEYDWHVFSIVTSKFPGYQDF
IAILKTTVDNSFVGWDLQHIITLDAVDGDSRTQIQLKKVQSPVVLLYCSKDEAAYILEEA
RSLGLTGFGYIWIVPSLTTGNPEITPDTFPSGMISVSYDDWDYPLEARVRDGLGIITTAA
AAMLEEFGDIPEAKTSCYGQMEKTKLPPSALHKYMMNVTWDGKDLSFTEDGYQVHPKLVV
IVLNKDREWEKMGKWENQSLTLKYPVWPRFNSFGDAETDDNHLTIVTLEEKPFVIVENVE
RLTGTCMRNSVPCRKHIKDNTTEGGGTYIKKCCKGFCIDILKKIARNVKFTYDLYLVTNG
KHGKKINNVWNGMVGEVVNKKAVMAVGSLTINEERSEVIDFSVPFVETGISVMVSRSNGT
VSPSAFLEPFSASVWVMMFVMLLLVTAMAVFMFEYISPLGFNRNLAQGKDPHGPSFTMGK
AVWLLWGLVFNNSVPVQNPKGTTSKFIVSVWAFFAVIFLASYTANLAAFMIQEEFVDQVT
GLSDKKFQSPYSYSPPFRFGTVPNGSTERNIRKNYPDMHQYMVKYHQTGVNDALVSLKTG
KLDAFIYDAAVLNYMAGRDEGCKLVTIGSGYIFATTGYGIALQKGSLWKRPVDLAILAII
GDGEMEELEAQWLTGICHNEKNEVMSSQLDVDNMAGVFYMLATAMGLSLITFVWEHLFYW
RLRYCFTGVCSGTPGLLFSISRGIWSCIHGVHIDMKKKSSDLDFSPQSNMLKLIKSAKQM
TNMTNLSGSRINSPKRGGEFMHSASPMIMDMMAEKGNFIYTDNRSYAQKDIYGDSSDLQS
YLTNRHKDHLNNYIFQGQHPLTLNESNPNTVEVAVSADTAQTNAKPRALWKKSVDTLRQT
PQGPETLDPRLSMKSQRYLPEDTAHSDISDCSSRATSYKDPDNNKHLKPKDNLKKRSMTS
KYPRDCSEVELSYLKNKQGGAGREKIYTIDSDRELSLHSDPQHYREGRGLTTDDLDFGEI
YSDHNDNYRKCDQPIIHLNSSPMHQSESDLLPDNVYNKHYSLKEKNISPHESNDRFKQTH
CRSCLSKVPNYNTGPYAGARSPYNRCEACLHMGNLYDISEDQMLQEAMISPSMHHDDMFS
HYWPQTDGPHVQRRNRLKLSRQHSFDNIMLEKPKEIDLNRPARSVSLKEKDRFLEDSPYA
NMFNIKADKLFGSRSMLFNRNLEESKRSKSLYPDHPSDNPFMQSIRDDTRLVHGRSSSDI
YKQLAPIKARNENNLRSSVKSTTSYCSRDGRIPNDMYISEHVMPYVANKTSAYSAPRVLN
SSQFSNRRVYKKIPSLESDV-
Mutant Protein Sequence
MKSSTMGFSRLVLLTLPALLTLISISHAAEKVPILNIAVILGQTRYISDRDIRALWGKDDP
IDEHWHRPGSHCPDFGFYFLSDFDTHPWDPWWLIHDHGG-
In Situ Probe Sequence
CCAGCTCTCCTTACCCTCATCAGCATCTCACATGCGGCAGAGAAAGTGCCCATACTGAACATTGCTGTGATCCTGGGTCAGACTCGCTACATCTCTGACAGAGATATACGGGCTCTGTGGAGCAAAGATGATCCTATTGATGTCAATGTGGTGACCCTGCTGGTGAATGAAACAGACCCCAAAAGCATCATTACCCATGTGTGTGACCTCATGTCCGGGACTAAAATCCACGGGGTGGTGTTCGGGGACGGCACAGACCAGGAAGCCATTGCCCAGATTTTGGATTTTATTTCCTCTCAGACTTTGATACCCATCCTTGGGATCCATGGTGGCTCATCCATGATCATGGCGGATAAGGATGTAAAATCAACATTCTTCCAGTTTGGGGCCTCCATCCAGCAGGAGGCGCTGCTCATGCTCAACATCATGGAAGAGTACGACTGGCACGTCTTCTCCATAGTGACCAGCAAGTTTCCAGGCTACCAGGATTTCATCGCTATCCTGAAGACCACGGTGGACAACAGCTTCGTGGGTTGGGACCTCCAGCACATCATCACCCTGGACGCTGTGGATGGAGACTCCAGGACCCAGATCCAGCTGAAAAAAGTTCAATCTCCTGTCGTCCTCTTGTACTGCTCTAAGGATGAAGCGGCGTACATCCTGGAGGAGGCCCGGTCACTGGGTCTGACCGGGTTCGGGTACATCTGGATCGTGCCAAGCTTGACCACAGGTAACCCGGAGATCACTCCAGATACGTTCCCCTCTGGAATGATCTCCGTGTCCTACGATGACTGGGACTATCCTCTTGAGGCACGGGTTCGAGATGGGCTTGGGATTATAACCACTGCAGCTGCGGCCATGTTGGMutant Genotyping Size
423Ribosome Profiling Development
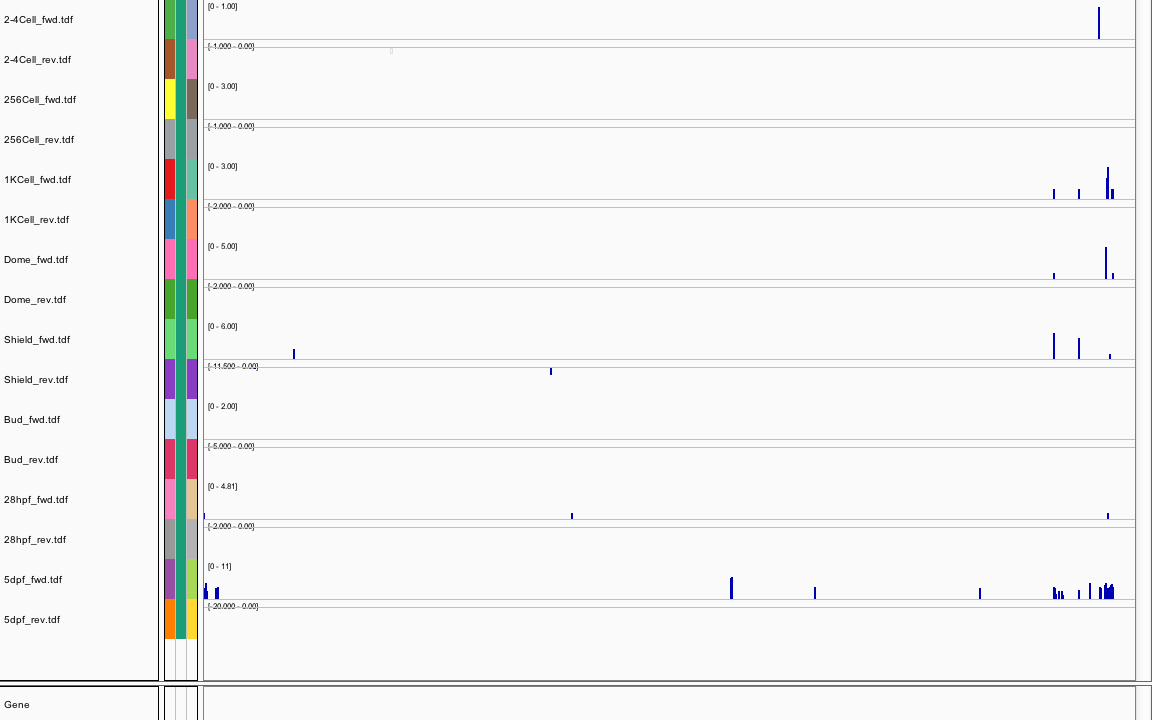
Sequence 2
Ensembl Gene Name
grin2abHarvard Allele
a279Mutation Area - WT DNA Sequence
GATGCTTGCTATAATGGAGGAGTATGACTGGCATGTCTTCTCGGTGGTCACCACCAAGTTCCCTGGTTTCCAGGAGTTCATAGCCACTGTGCGCGTGACTGTGGACCACAGCTTTGTTCGTTGGGACCTTCAGAGCGTAGTGACTCTAGACGGCGTCGGTGAACCCGAGGACCGAGCTCATGTGCAGCTCAAACGCATCCAGTCGCCCGTCATCCTCCTCTACTGTTCCAAAGATGAAGCAGCGCTGGTTATGGAGGAAGCCCGTTCGCTAGGGCTCACAGGGGCTGGCTTTGTGTGGATTGTTCCTAGCCTGACGACAGGGAATCTCGAACATATCCCGGAAGCCTTTCCGGCAGGGATGATATCTGTGACCTATGATGAATGGGATTATCCCCTGGAGACTCGCCTGCAGGATGCTGTGGGAATTATCAGTTCGGCAGCCGCTGCCATGTTCAGAGAAGAAGGCCGGATACCTGATGGAACAGCCAGCTGCTACGGCCAATCAGAGAAGCCGGAGGTGCMutation Area - Mutant DNA Sequence
GATGCTTGCTATAATGGAGGAGTATGACTGGCATGTCTTCTCGGTGGTCACCACCAAGTTCCCTGGTTTCCAGGAGTTCATAGCCACTGTGCGCGTGACTGTGGACCACAGCTTTGTTCGTTGGGACCTTCAGAGCGTAGTGACTCTAGACGGCGTCGGTGAACCCGAGGACCGAGCTCATGTGCAGCTCAAACGCATCCAGTCGCCCGTCATCCTCCTGTTCCAAAGATGAAGCAGCGCTGGTTATGGAGGAAGCCCGTTCGCTAGGGCTCACAGGGGCTGGCTTTGTGTGGATTGTTCCTAGCCTGACGACAGGGAATCTCGAACATATCCCGGAAGCCTTTCCGGCAGGGATGATATCTGTGACCTATGATGAATGGGATTATCCCCTGGAGACTCGCCTGCAGGATGCTGTGGGAATTATCAGTTCGGCAGCCGCTGCCATGTTCAGAGAAGAAGGCCGGATACCTGATGGAACAGCCAGCTGCTACGGCCAATCAGAGAAGCCGGAGGTGCGuide RNA Target Sites
TTCTTCTCTGAACATGGCAGCGG
ATTGTTCCTAGCCTGACGACAGG
CATCTTTGGAACAGTAGAGGAGG
CCCCTGTGAGCCCTAGCGAACGG
Genotyping Primers
f, TCATGTGCAGCTCAAACGCATCC
r, AGGAACAATCCACACAAAGCCAGC
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
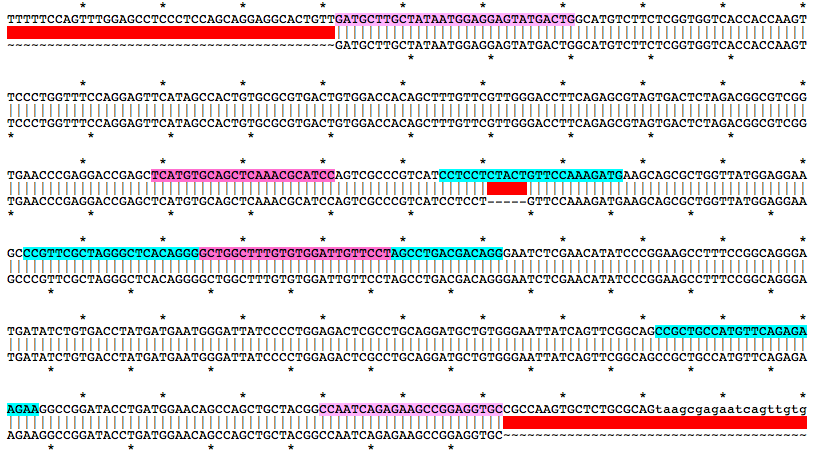

Allele
5bpDWT Genotyping Size
130WT Protein Sequence
MMGAVVRLMVLWALLGGSQAAEKPPSLSIAVILGQTPTVSESALRPARGIGAPLDVTVLS
IQVNQTDPGSMLTHLCELMSRQRLHGLVYGDGTDQEAVAQMLDFVSSQTSMPILGIHGGS
AMTMAEKDSSSSFFQFGASLQQEALLMLAIMEEYDWHVFSVVTTKFPGFQEFIATVRVTV
DHSFVRWDLQSVVTLDGVGEPEDRAHVQLKRIQSPVILLYCSKDEAALVMEEARSLGLTG
AGFVWIVPSLTTGNLEHIPEAFPAGMISVTYDEWDYPLETRLQDAVGIISSAAAAMFREE
GRIPDGTASCYGQSEKPEVPPSALRRYMMGVNLGGKDHSFMDDGYQVNPKLVVIVLNTER
EWEKMGRWENHSLRLKFPVWPRYNSFGDMDADENHLSIVTLEERPFVIVDDVDILTGTCM
RNSVPCRKHIKDNTTEGSYVKKCCKGFCIDILKKIARNVKFTYDLYLVTNGKHGKKINNV
WNGMVGEVVYKKAVMAVGSLTINEERSEAIDFSVPFVETGISVMVSRSNGTVSPSAFLEP
FSASVWVMMFVMLLIVTAIAVFLFEFISPLGFNRNLAQGKDPHGPSFTIGKAVWLLWGLV
FNNSVPVQNPRGTTSKFIVSVWAFFAVIFLASYTANLAAFMIQEEFVDQVTGLSDRKFQS
PYSYSPPFRFGTVPNGSTERNIRKNYPAMHQYMVKYHQTGVVDALVSLKTGKLDAFIYDA
AVLNYMAGRDDGCKLVTIGSGYIFATTGYGIAMQKGSYWKRLVDLAILGIIGDGEMEELE
AQWLTGICHNEKNEVMSSQLDVDNMAGVFYMLAAAMALSLITFVWEHLFYWRLRYCFTGL
CSGRPGILFTISRGIWSCIHGVHIELKKAPELDFSPQANMLHLLKSAKEITTLGVTSPKR
PMSSILPPTSGLLELMGGGKGNNLAPKNPPYSNTSTMLSNRLKEPLNNTLLPGQRALSLT
DAQPGVAGNGGSTGGGETMRPPASKPRALWKKSVETLRQNQGPEDSSPPSAPQAAETLPM
KSQRYLPEEAPSIDISDCSARVDLQKQHKLKETIRKRNRLPRDLSDVELSTLRSQPSQQS
NHSAHIYSQSSQGSQIYSIDSIDQEHGLHYPDIFSDHRDNKHRPPSSSEQQPILQLSSSL
LSQAHEPDVMQTAVAKDKKYNTYGKPGGAGSIAGGPASTQDPRQTHCRSCLSKVSSYSGL
YTVRSPQYRCDACLHSGNLYDISEDQFLHPDGRGTGGGSVDRAFSPFLPQPDASFTAYLP
QSDQEGDGLISHYQNEGVIIRQHSYENLPQKSSSRRVSLQDNTPYANIPSATLMPDKPPK
TQPVHLNHTLSGVGVSGLGIGRRSKSMYPARPTENPFLQTLRDDQRLAHGRSSTDIYKQL
VPLTLGQADGSVLSMGMPTPDLYYSEHVMPYVAPPPIGTYTAPRALSAGGARRVYKRMPS
IESDV-
Mutant Protein Sequence
MMGAVVRLMVLWALLGGSQAAEKPPSLSIAVILGQTPTVSESALRPARGIGAPLDVTVLS
IQVNQTDPGSMLTHLCELMSRQRLHGLVYGDGTDQEAVAQMLDFVSSQTSMPILGIHGGS
AMTMAEKDSSSSFFQFGASLQQEALLMLAIMEEYDWHVFSVVTTKFPGFQEFIATVRVTVD
HSFVRWDLQSVVTLDGVGEPEDRAHVQLKRIQSPVILLFQR-
In Situ Probe Sequence
CCACAGTGCTCACATCTACTCTCAGAGCAGCCAGGGGAGTCAGATTTACTCCATTGACTCTATTGACCAGGAGCACGGGCTGCACTATCCTGATATCTTCAGTGATCACCGTGACAACAAGCATCGTCCACCCTCTTCCTCAGAGCAGCAGCCAATCCTGCAGCTCAGCAGCAGTCTCCTCTCGCAGGCCCATGAACCAGACGTGATGCAAACCGCTGTCGCTAAGGATAAAAAATACAACACGTATGGAAAGCCCGGAGGGGCGGGAAGCATTGCCGGCGGGCCTGCTTCTACTCAGGATCCACGACAGACACACTGCCGTAGCTGCTTGTCGAAAGTGTCCAGCTATTCTGGGCTTTACACTGTACGGTCCCCTCAGTACCGCTGCGATGCCTGCCTGCACTCGGGCAACCTGTATGACATCAGTGAAGATCAGTTTCTCCATCCAGATGGTCGGGGCACTGGAGGTGGAAGTGTTGACAGGGCCTTCTCTCCTTTTCTTCCCCAGCCTGATGCTTCCTTCACAGCCTATCTTCCCCAGAGTGACCAAGAGGGCGACGGTCTCATTAGTCACTACCAGAATGAGGGCGTAATCATCCGCCAGCACTCGTACGAGAACCTGCCACAGAAGAGCTCCTCTCGGCGGGTAAGCCTGCAAGACAACACGCCGTATGCTAACATTCCCTCGGCGACTCTAATGCCGGACAAGCCCCCGAAAACTCAGCCGGTGCACCTTAACCACACACTCAGTGGAGTGGGCGTCAGCGGGCTCGGCATCGGCCGTCGCAGCAAGTCGATGTATCCTGCGCGCCCAACGGAAAACCCCTTTCTCMutant Genotyping Size
125Ribosome Profiling Development
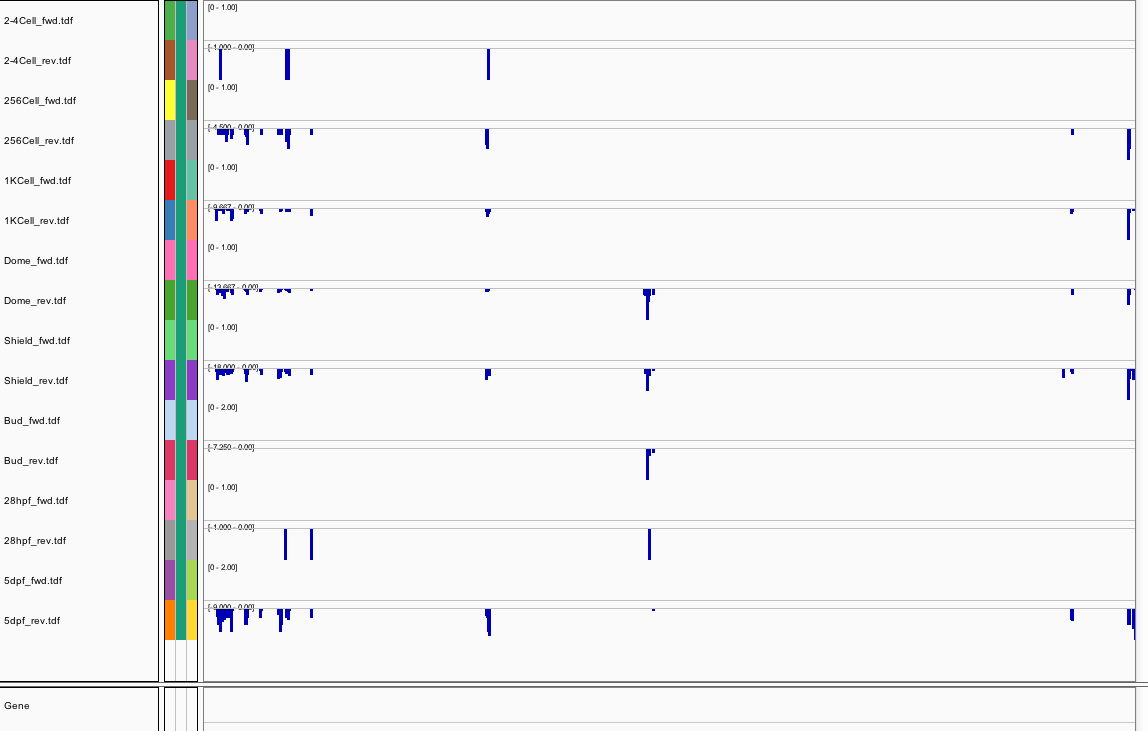
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = 0.0142 23 hethom vs 17 homhom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
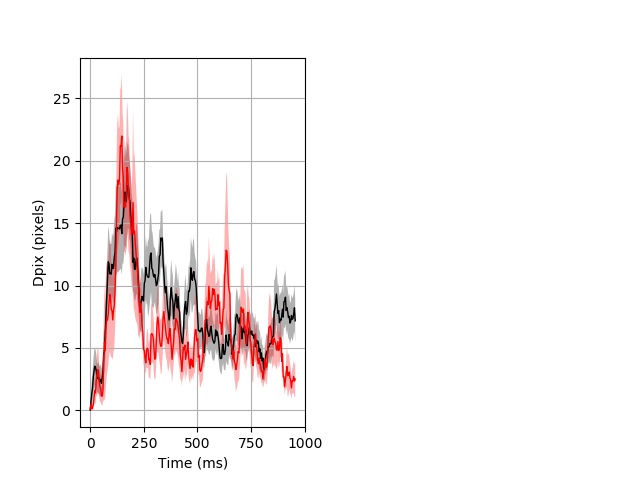
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0155 23 hethom vs 17 homhom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
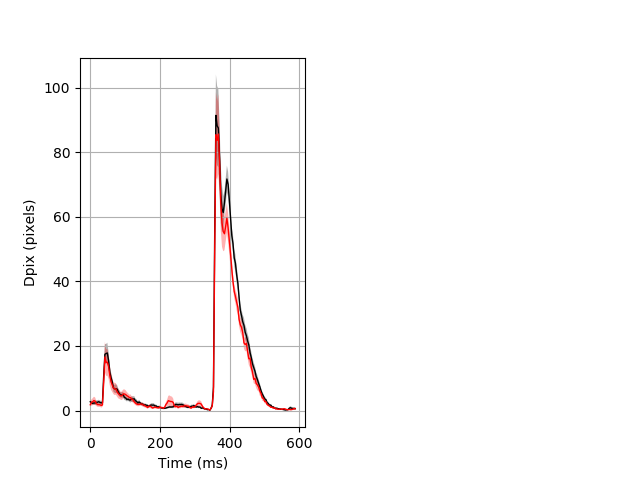
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = 0.0172 Merged section Pvalue = 0.0148 24 hethet vs 17 homhom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
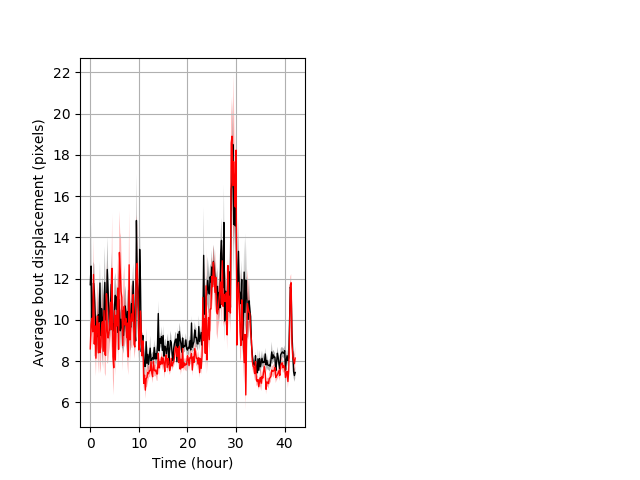
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 24 hethet vs 17 homhom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
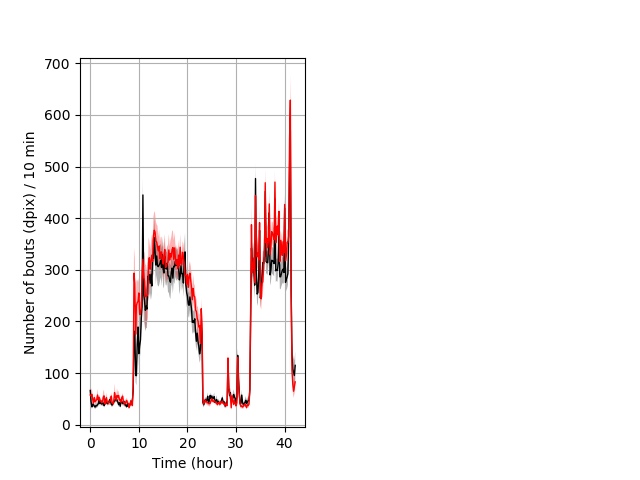
Behavior Data Description 5
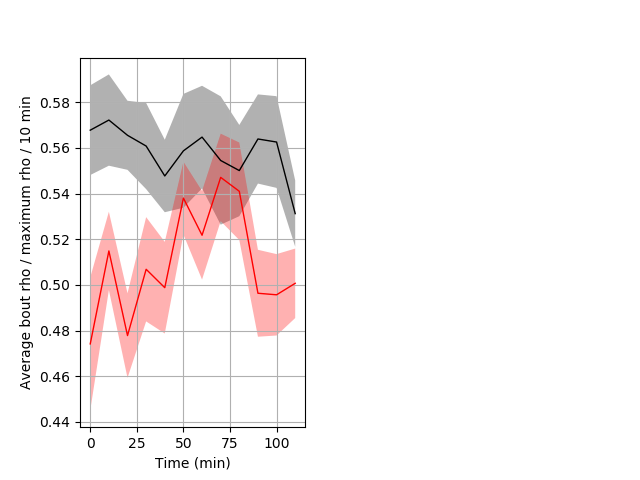
Location in well entire protocol Graph Pvalue = 0.0088 Merged section Pvalue = 0.0119 24 hethet vs 17 homhom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
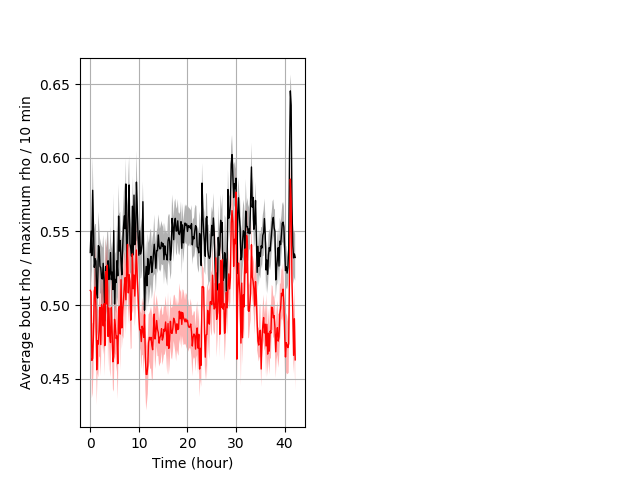
Behavior Data Description 6
Dark flash Graph Pvalue = 0.0102 24 hethet vs 17 homhom boxgraph_pd__ribgraph_mean_ribbon_velocityresponse_dist_ddarkflash103.pngBehavior Data Graph 6
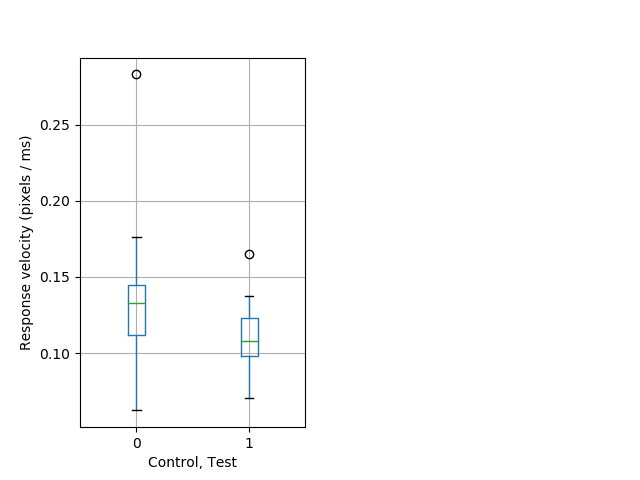
Behavior Data Description 7
Features of movement Graph Pvalue = 0.001 22 hethet vs 27 hethom ribgraph_mean_ribbonbout_avebouttime_min_day2night1.pngBehavior Data Graph 7
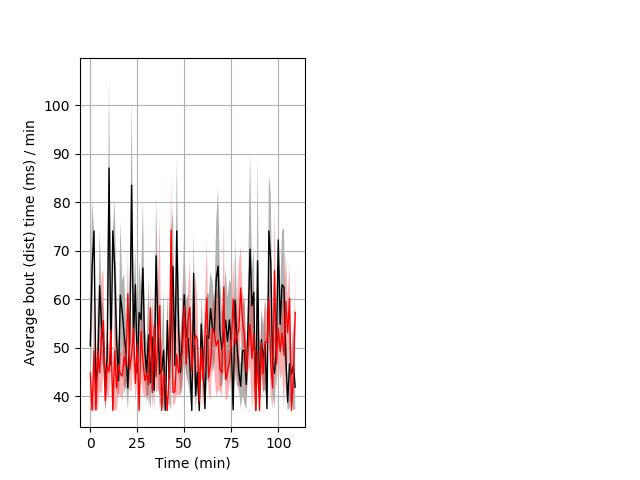
Behavior Data Description 8
Location in well Graph Pvalue = 0.001 18 homhet vs 17 homhom ribgraph_mean_ribbonbout_averhofrac_10min_day2night2.pngBehavior Data Graph 8
Behavior Data Description 9
Location in well Graph Pvalue = 0.004 24 hethet vs 17 homhom ribgraph_mean_ribbonbout_interboutaverhofrac_min_day1taps.pngBehavior Data Graph 9
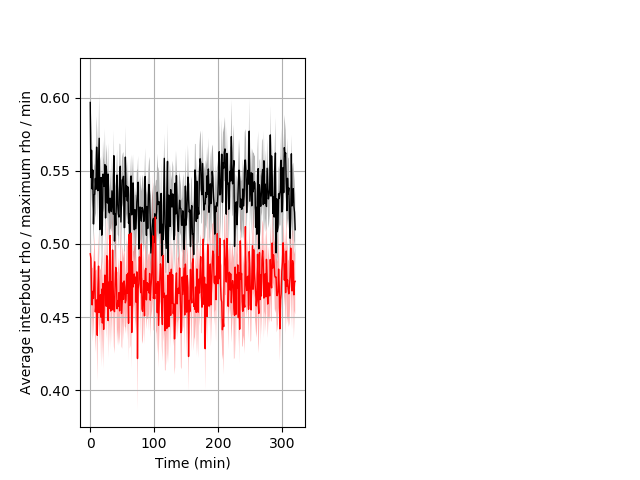
Behavior Data Description 10
PPI tap Graph Pvalue = 0.00462 22 hethet vs 22 homhom boxgraph_pd__ribgraph_mean_ribbon_displacement_dist_shortnightprepulseinhibition100b.pngBehavior Data Graph 10
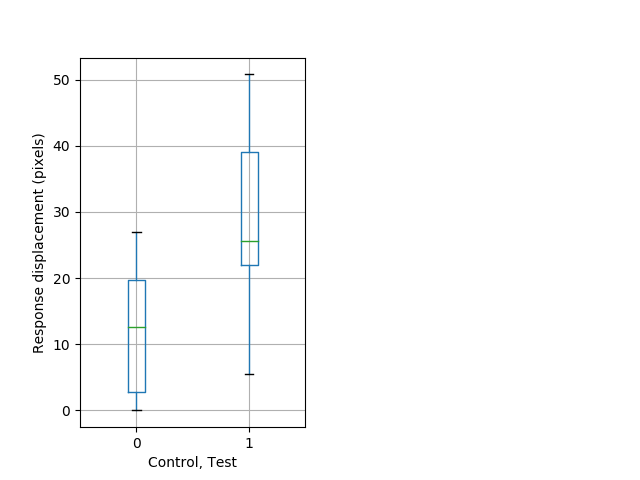
Behavior Data Description 11
Strong tap Graph Pvalue = 0.000804 23 hethom vs 17 homhom boxgraph_pd__bigmovesribgraph_mean_ribbon_totaldistanceresponse_dist_shortdayprepulseinhibition102.pngBehavior Data Graph 11
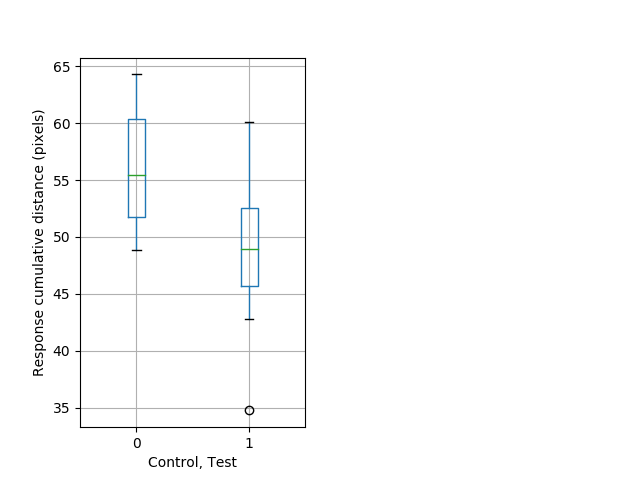
Behavior Data Description 12
Tap habituation Graph Pvalue = 0.00486 18 homhet vs 17 homhom boxgraph_pd__ribgraph_mean_ribbon_freqresponse_dpix_nighttaphab102.pngBehavior Data Graph 12
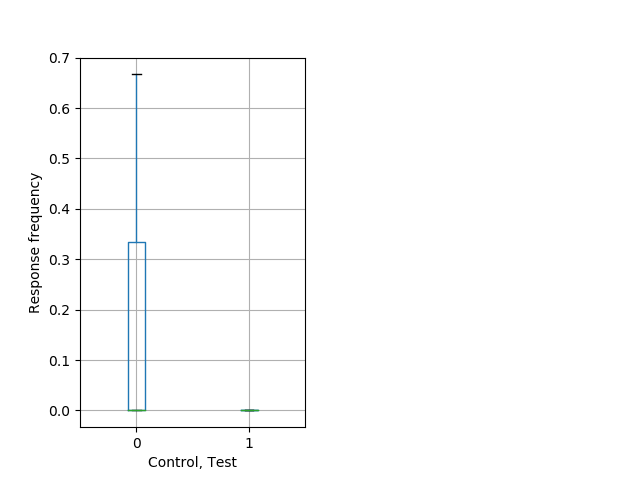
Behavior Data Description 13
Weak tap Graph Pvalue = 0.00704 23 hethom vs 17 homhom boxgraph_pd__bigmovesribgraph_mean_ribbon_latencyresponse_dpix_dayprepulseinhibition100a.pngBehavior Data Graph 13
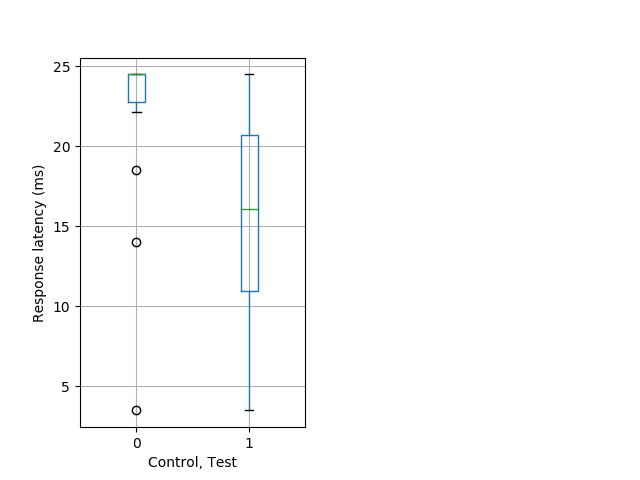
Behavior Data Description 14
NoneBehavior Data Graph 14
None