Locus Rank
64Gene
Id
216Gene Name
AKT3Duplicated
TrueMaternal
TrueGene Not Within Locus (Nearby
FalseGene Description
Mouse Phenotype
Additional Image
Allen Brain
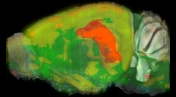
Allen Regions
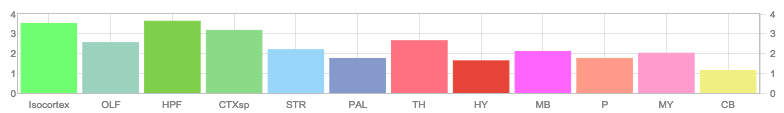
Zfin In Situ
Zfin Brain Areas
Published Zebrafish Pehnotype
Not In Allen Brain
FalseZFIN Link
http://zfin.org/search?q=akt3&fq=category%3A%22Gene+%2F+Transcript%22&category=Gene+%2F+Transcript&fq=type%3A%22Gene%22Allen Link
http://mouse.brain-map.org/gene/show/23550More Additional Images
Papers
Locus Rank
Locus Rank
64Genes in Region
AKT3 SDCCAG8Associated Snps
rs77149735 chr1_243881945_I rs10803138 rs14403GWAS Region (hg19)
chr1:243503719-244002945Genes Skipping
Ricopili Plot Surrounding Area
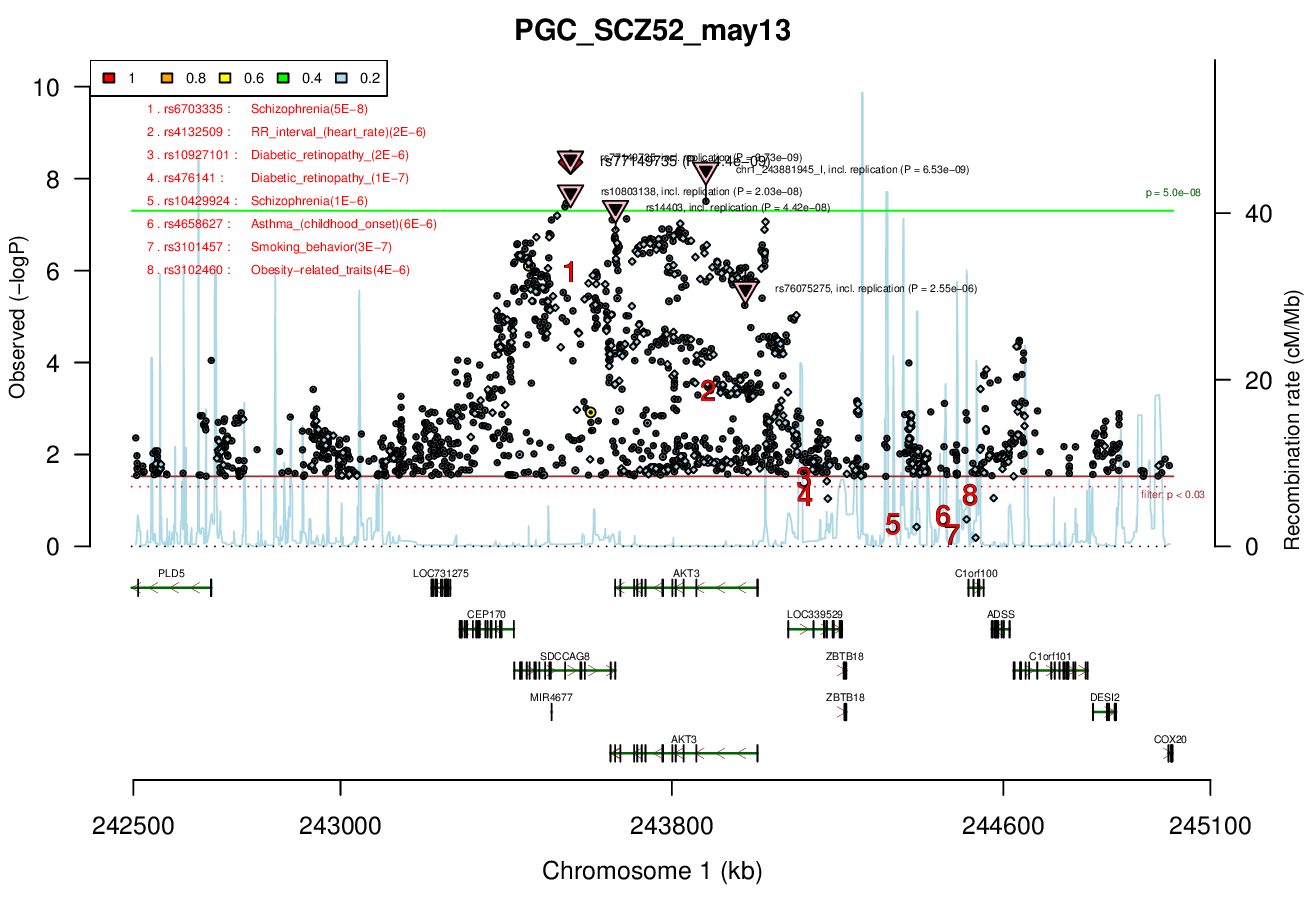
Distance On Each Side Of Locus In Above Plot
1 MBRicopili Plot
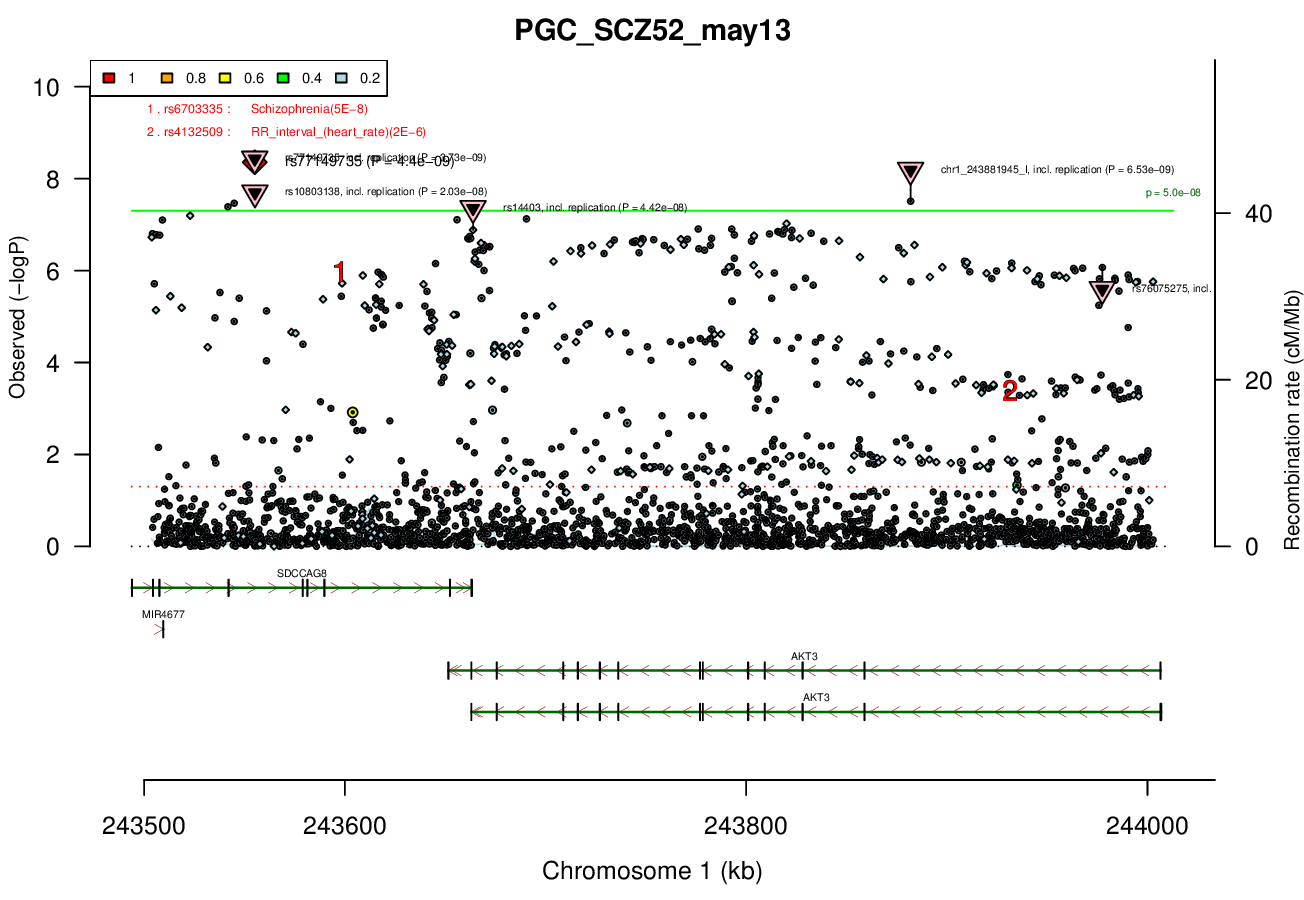
Gwas And Surrounding Region Pic
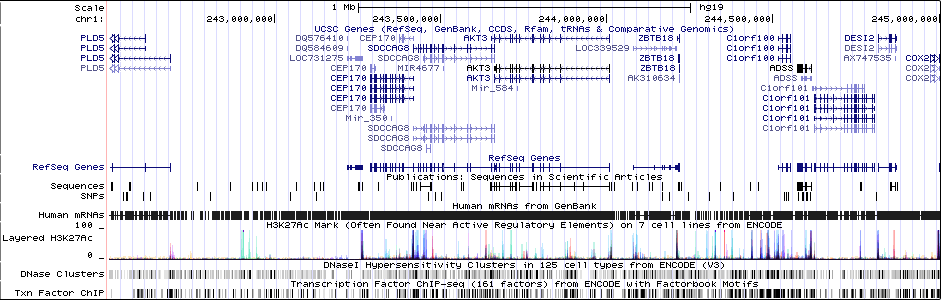
Gwas Region Pic
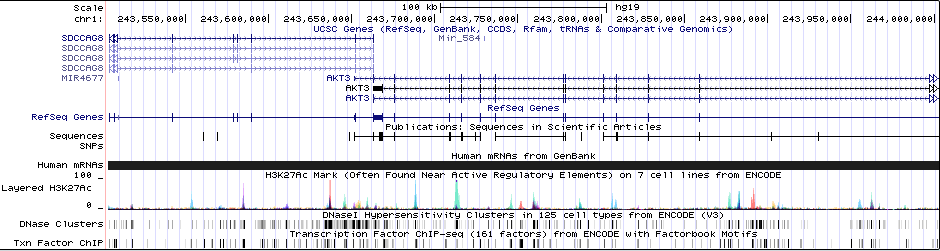
Sequence 1
Ensembl Gene Name
akt3aHarvard Allele
a229Mutation Area - WT DNA Sequence
tcctagaaacatgggctttgtgtgtgtttccaaagGGATGAGTGGGCAGAGGCGATCCAGATGGTGGCAGATAAACTCCAAAGACAGGAAGAGGAGCGAATCCAGTGTAGTCCCACCTCACAGATAGACAACATGGGAGAGGAGGAGATGGACACGTCTATCAGCCACCACAAACGCAAgtgcctgcacacactgatgtgggagttcMutation Area - Mutant DNA Sequence
ttcctagaaacatgggctttgtgtgtgtttccaaagGGATGAGTGGGCAGAGGCGATCCGAGGCGATCAGAGGCGAGATGGACACGTCTATCAGCCACCACAAACGCAAGgtgcctgcacacactgatgtgggagttcGuide RNA Target Sites
GGCAGAGGCGATCCAGATGGTGG
TAAACTCCAAAGACAGGAAGAGG
ATGTTGTCTATCTGTGAGGTGGG
CAACATGGGAGAGGAGGAGATGG
Genotyping Primers
f, tcctagaaacatgggctttgtgtgtgtttcc
r, gaactcccacatcagtgtgtgcag
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
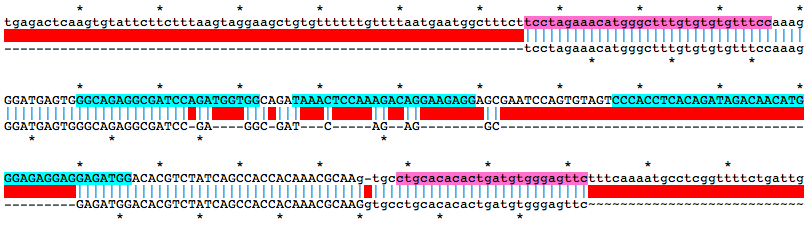
Allele
71bpDWT Genotyping Size
207WT Protein Sequence
MSDVTVVKEGWVQKRGEYIKNWRPRYFLLKTDGSFIGYKEKPQDADLPYPLNNFSVAKCQ
LMKTERPKPNTFIIRCLQWTTVIERTFHVDTPEERDEWAEAIQMVADKLQRQEEERIQCS
PTSQIDNMGEEEMDTSISHHKRKTMNDFDYLKLLGKGTFGKVILVREKASGKYYAMKILK
KEVIIAKDEVAHTLTESRVLKNTRHPFLTSLKYSFQTKDRLCFVMEYVNGGELFFHLSRE
RVFSEDRTRFYGAEIVSALDYLHSAKIVYRDLKLENLMLDKDGHIKITDFGLCKEGITDA
ATMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHEKLFELILM
EDIKFPRTLSADAKSLLSGLLIKDPNKRLGGGPDDAKEIMRHSFFTGIDWQDVYDKKLIP
PFKPQVSSETDTRYFDEEFTAQTITITPPEKFDEDGMDCMDNERRPHFPQFSYSASGRE-
Mutant Protein Sequence
MSDVTVVKEGWVQKRGEYIKNWRPRYFLLKTDGSFIGYKEKPQDADLPYPLNNFSVAKCQ
LMKTERPKPNTFIIRCLQWTTVIERTFHVDTPEERDEWAEAIRGDQRRDGHVYQPPQTQDNE-
Mutant Genotyping Size
138Transcript Plot
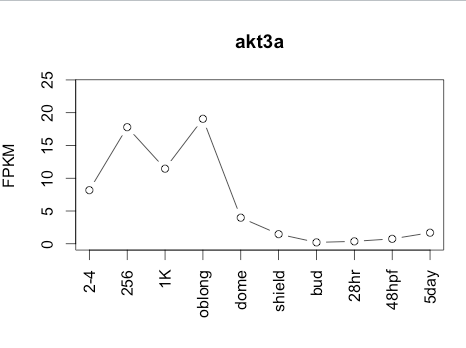
Sequence 2
Ensembl Gene Name
akt3bHarvard Allele
a230Mutation Area - WT DNA Sequence
tttggtcagAGTGTCAGCTGATGAAGACTGAGAGACCGAAACCAAACACCTTCATCATCAGGTGTCTGCAGTGGACCACTGTGATCGAGCGCACCTTCCATGTCGACACGCCAGAGGAGAGgtaaaacacgctaaaacacgcaagctagccaaccaatgaggatgcagtcaacaactgaataacatttacagggcatggaaatccttgatgattgtatctagtgtattgaaggtttaaatagtgcatatattgtggatgaagtgcttgaaaatgtaattgttgcttttataataatttatcctagtaaatagttaataaaaatgatatattcaataattaattagttgttttgtattgagtgaaaagcacatacatgctaaatacataaatgctttgaacacataatagtaaacatgtatatataacagatgaatgataaataaataacatcacttaatgataataataataataataataataatgtttggactgtggggtaaatgggaacacccggagaaaacccacaacaacactgggagaacatgcaaactgacccagctggtactcgcaccagtgaccttcttgcttagaggcaacggtgctaaccactaagccaccgtgtcgccctctgatgcatatcggctaattgaattattctttttttttttgaacCAGGGACGAGTGGGTCGAGGCCATTCAGATGGTGGCAGACAAGCTGGCCAAACAGGAGGAAGAAGGAATCCTGTGTAGCCCAATTTCTCAGATTGAAAACGTCAACGAGGAGGAGATGGACACGTCAACCAGTCATCACAAACGAAAGgtagactctctgccaggagcagaagcgttgtaccttgaggttcacaggMutation Area - Mutant DNA Sequence
tttggtcagAGTGTCAGCTGATGAAGACTGAGAGACCGAAACCAAACACCTTCATCATCAGGTGGACCACTGTGATCGAGCGCACCTTCCATGTCGACACGCCAGAGGAGAGgtaaaacacgctaaaacacgcaagctagccaaccaatgaggatgcagtcaacaactgaataacatttacagggcatggaaatccttgatgattgtatctagtgtattgaaggtttaaatagtgcatatattgtggatgaagtgcttgaaaatgtaattgttgcttttataataatttatcctagtaaatagttaataaaaatgatatattcaataattaattagttgttttgtattgagtgaaaagcacatacatgctaaatacataaatgctttgaacacataatagtaaacatgtatatataacagatgaatgataaataaataacatcacttaatgataataataataataataataataatgtttggactgtggggtaaatgggaacacccggagaaaacccacaacaacactgggagaacatgcaaactgacccagctggtactcgcaccagtgaccttcttgcttagaggcaacggtgctaaccactaagccaccgtgtcgccctctgatgcatatcggctaattgaattattctttttttttttgaacCAGGGACGAGTGGGTCGAGGCCATTCAGAAGGAATCCTGTGTAGCCCAATTTCTCAGATTGAAAACGTCAACGAGGAGGAGATGGACACGTCAACCAGTCATCACAAACGAAAGgtagactctctgccaggagcagaagcgttgtaccttgaggttcacaggGuide RNA Target Sites
TCATCATCAGGTGTCTGCAGTGG
ATGTCGACACGCCAGAGGAGAGg
GGTCGAGGCCATTCAGATGGTGG
CTGGCCAAACAGGAGGAAGAAGG
Genotyping Primers
f, agtgaccttcttgcttagaggcaacg
r, cctgtgaacctcaaggtacaacgc
Alignment of WT and Mutant Sequences w/ gRNA Targets (cyan) and Genotyping Primers (pink) Shown on WT Sequence
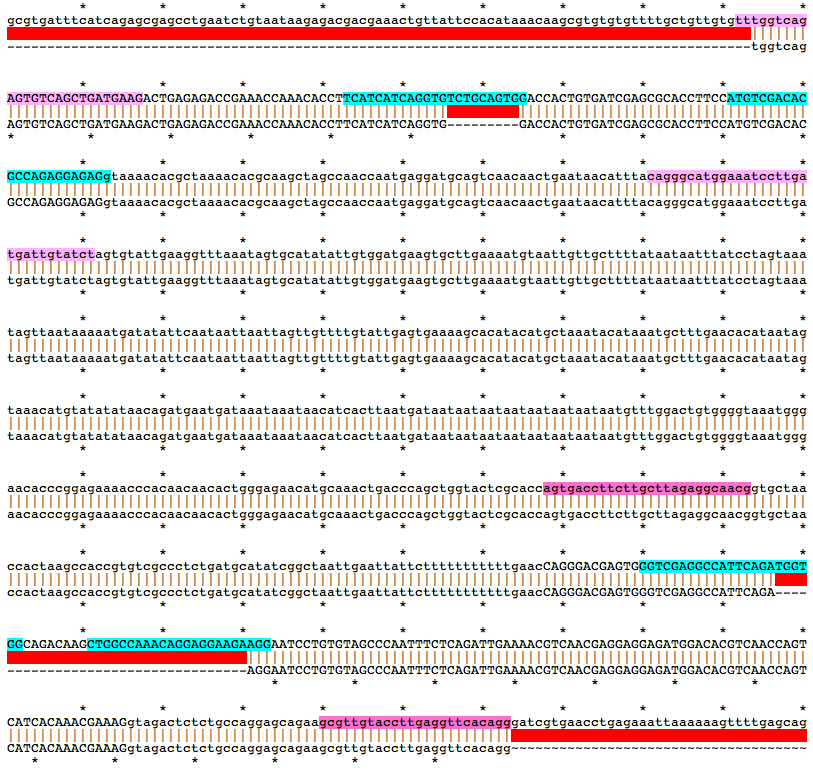

Allele
9bpD,34bpDWT Genotyping Size
296WT Protein Sequence
MNDLNVVKEGWVQKRGEYIKNWRPRYFLLKTDGSFIGYKEKPQDADLAYPLNNFSVAKCQ
LMKTERPKPNTFIIRCLQWTTVIERTFHVDTPEERDEWVEAIQMVADKLAKQEEEGILCS
PISQIENVNEEEMDTSTSHHKRKTMNDFDYLKLLGKGTFGKVILVREKASGTYYAMKILK
KEVIIAKDEVAHTLTESRVLKNTRHPFLTSLKYSFQTKDRLCFVMEYVNGGELFFHLSRE
RVFSEDRTRFYGAEIVSALDYLHSAKIVYRDLKLENLMLDKDGHIKITDFGLCKEGITDA
ATMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHEKLFELILM
EEIKFPRTLSADAKSLLSGLLIKDPNKRLGGGPDDAKEIMRHSFFTALDWQDVYDKKLVP
PFMPQVSSETDTRYFDEEFTAQTITITPPEKYDEDGMDAADSERRPHFPQFSYSASGRE-
Mutant Protein Sequence
MNDLNVVKEGWVQKRGEYIKNWRPRYFLLKTDGSFIGYKEKPQDADLAYPLNNFSVAK
CQLMKTERPKPNTFIIRWTTVIERTFHVDTPEESRDEWVEAIQKESCVAQFLRLKTSTRR
RWTRQPVITNERQ-
In Situ Probe Sequence
AAGCCAGCGGCACGTATTATGCAATGAAGATCCTGAAAAAGGAGGTTATTATTGCCAAGGATGAAGTTGCTCACACACTCACGGAGAGCAGAGTTTTGAAGAACACTAGGCACCCATTTCTAACTTCACTGAAGTATTCCTTTCAGACGAAGGATCGATTATGTTTTGTTATGGAGTATGTGAATGGAGGAGAGTTGTTTTTTCATTTGTCGAGAGAACGAGTGTTTTCGGAGGACAGAACCCGATTCTACGGCGCTGAGATCGTTTCCGCGTTAGATTACTTACACTCAGCGAAGATTGTTTACCGAGACCTGAAGCTGGAAAACCTGATGTTGGATAAAGACGGTCACATAAAGATCACTGATTTCGGCCTGTGTAAGGAGGGCATCACGGACGCCGCCACCATGAAGACGTTCTGTGGGACCCCGGAGTATTTAGCCCCTGAGGTATTGGAGGACAATGATTACGGTAGAGCCGTGGACTGGTGGGGTCTAGGAGTCGTCATGTATGAGATGATGTGTGGACGTCTGCCCTTTTATAACCAGGATCACGAGAAGCTGTTTGAGCTGATTCTGATGGAGGAGATCAAATTTCCCAGAACTCTCTCTGCTGATGCTAAGTCTTTACTGTCTGGCTTACTCATCAAGGACCCCAATAAGAGGCTCGGAGGAGGCCCGGATGATGCTAAAGAAATCATGAGACACAGTTTCTTCACGGCGCTGGACTGGCAGGACGTCTACGACAAAAAGCTGGTGCCGCCATTCATGCCCCAGGTCTCTTCAGAMutant Genotyping Size
262Ribosome Profiling Development
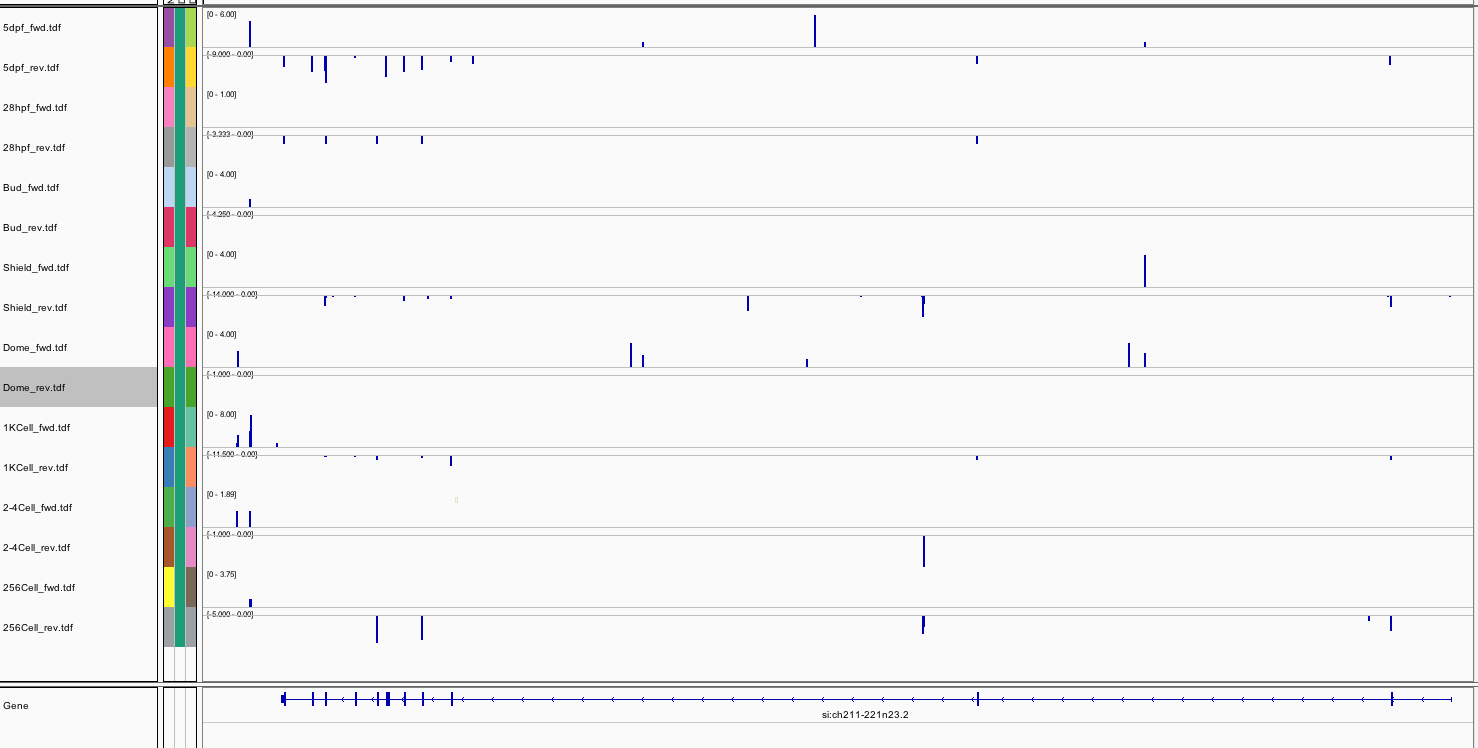
Behavior Data
Behavior Data Summary
NoneBehavior Data Description 1
Dark flash block 1 start Merged section Pvalue = 0.014 36 hetandwt vs 15 hom ribgraph_mean_ribbon_fullboutdata_dpix_a0darkflash103.pngBehavior Data Graph 1
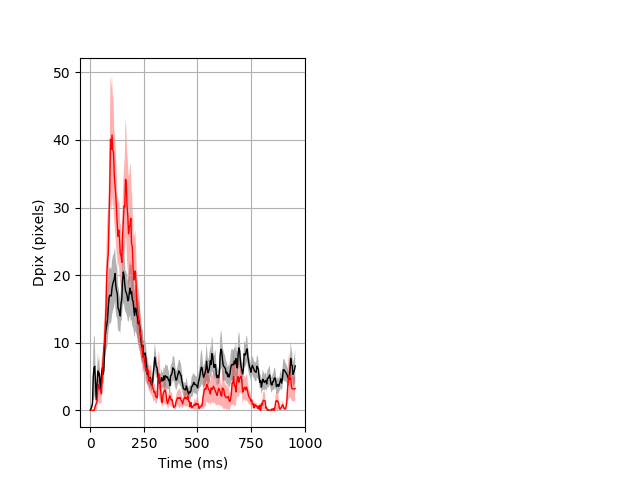
Behavior Data Description 2
Day all prepulse tap Merged section Pvalue = 0.0238 10 wt vs 10 hom ribgraph_mean_ribbon_fullboutdata_dpix_dayprepulseinhibition100d.pngBehavior Data Graph 2
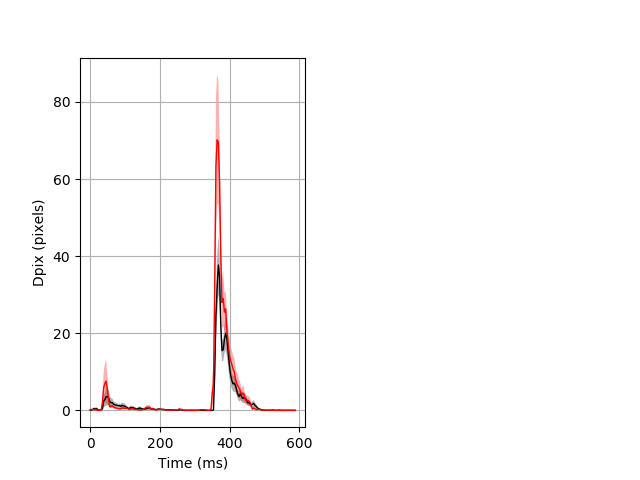
Behavior Data Description 3
Features of movement entire protocol Graph Pvalue = not significant Merged section Pvalue = 0.0255 22 wt vs 21 hom ribgraph_mean_ribbonbout_aveboutdisp_10min_combo.pngBehavior Data Graph 3
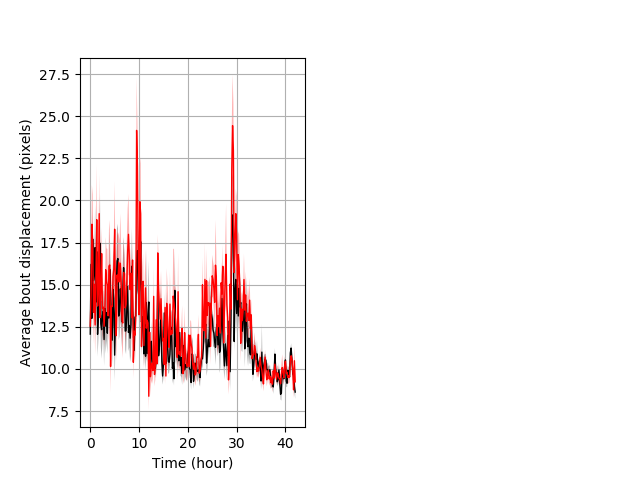
Behavior Data Description 4
Frequency of movement entire protocol Graph Pvalue = 2.63e-05 Merged section Pvalue = 0.00674 22 wt vs 21 hom ribgraph_mean_ribbonbout_dpixnumberofbouts_10min_combo.pngBehavior Data Graph 4
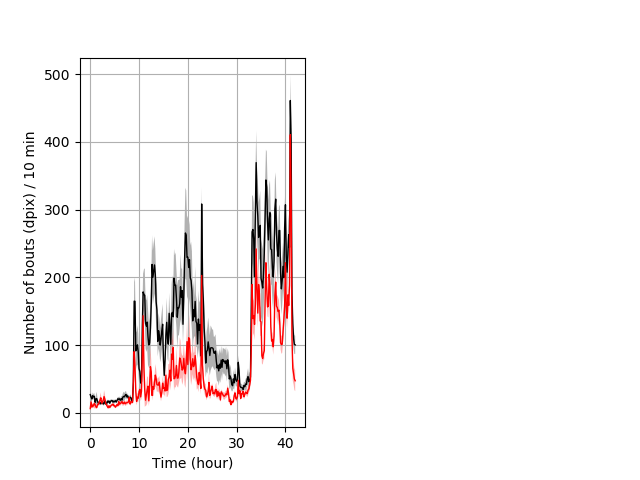
Behavior Data Description 5
Location in well entire protocol Graph Pvalue = not significant Merged section Pvalue = not significant 18 wt vs 16 hom ribgraph_mean_ribbonbout_averhofrac_10min_combo.pngBehavior Data Graph 5
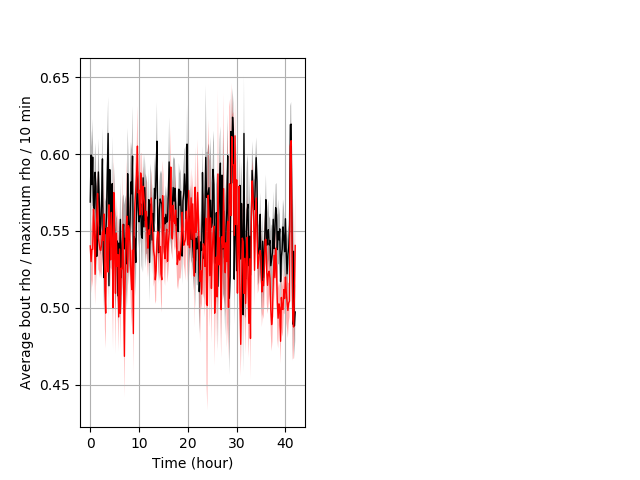
Behavior Data Description 6
Dark flash Graph Pvalue = 0.0107 36 hetandwt vs 15 hom boxgraph_pd__ribgraph_mean_ribbon_fullboutdatamaxloc_dpix_adarkflash103.pngBehavior Data Graph 6
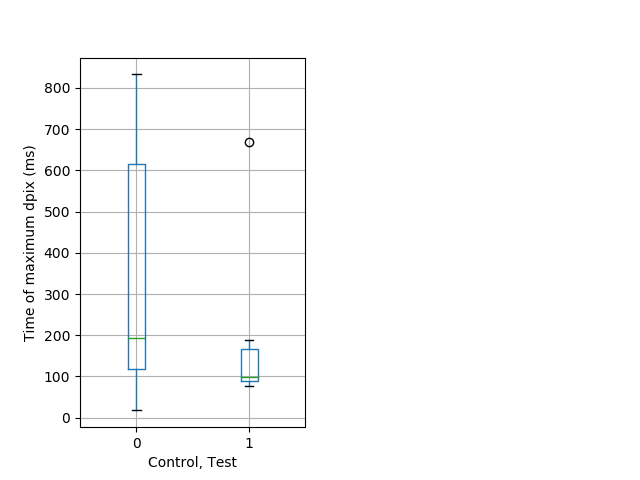
Behavior Data Description 7
Features of movement Graph Pvalue = 0.001 23 hetandwt vs 10 hom ribgraph_mean_ribbonbout_aveboutcumdpix_min_day1taps.pngBehavior Data Graph 7
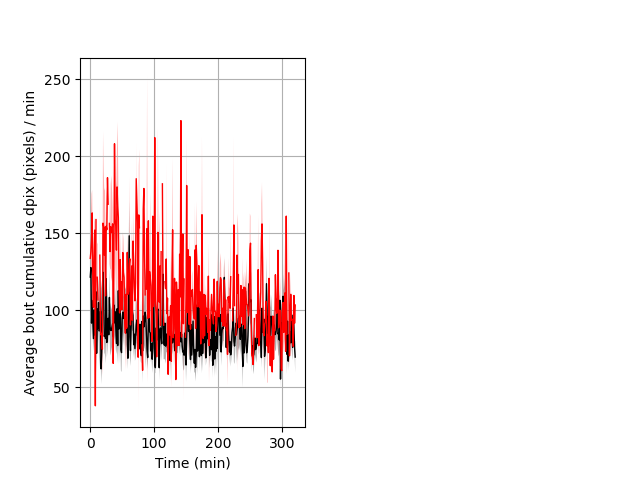
Behavior Data Description 8
Frequency of movement Graph Pvalue = 0.001 22 wt vs 21 hom ribgraph_mean_ribbon_dpixsecper_min_day2nightstim.pngBehavior Data Graph 8
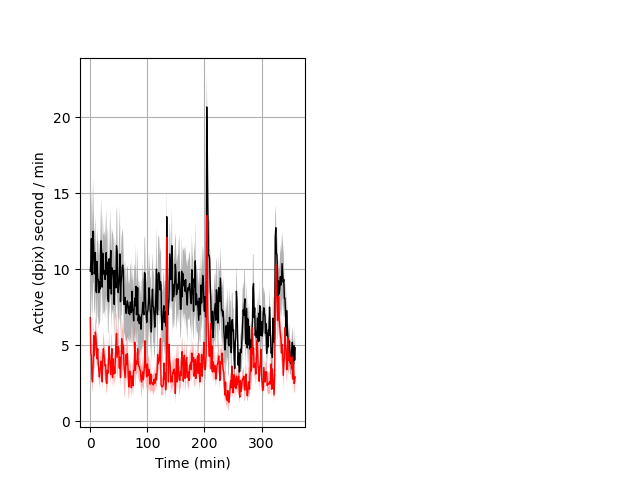
Behavior Data Description 9
PPI tap Graph Pvalue = 0.0039 23 het vs 15 hom boxgraph_pd__ribgraph_mean_ribbon_freqresponse_dpix_nightprepulseinhibition100b.pngBehavior Data Graph 9
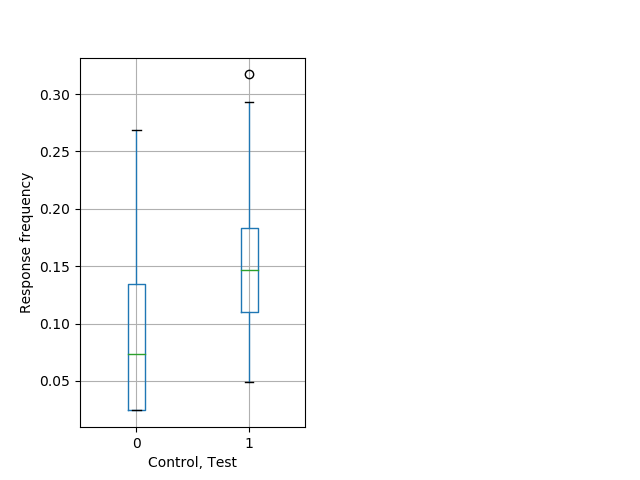
Behavior Data Description 10
Strong tap Graph Pvalue = 0.00162 61 hetandwt vs 16 hom boxgraph_pd__ribgraph_mean_ribbon_timeresponse_dpix_nighttappost102.pngBehavior Data Graph 10
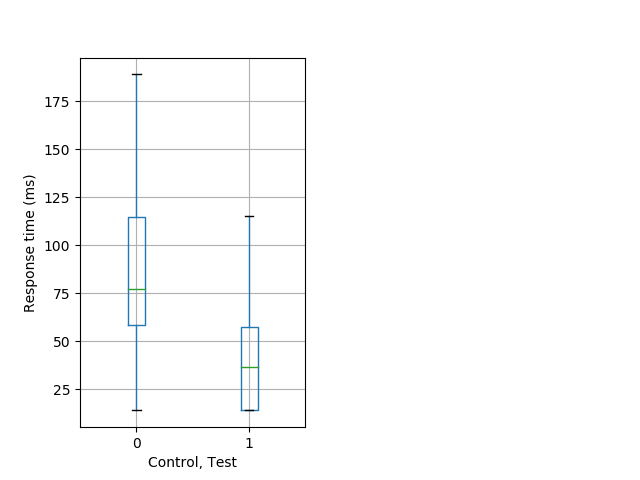
Behavior Data Description 11
Tap habituation Graph Pvalue = 0.000991 36 hetandwt vs 15 hom boxgraph_pd__smallmovesribgraph_mean_ribbon_speedresponse_dist_cdaytaphab102.pngBehavior Data Graph 11
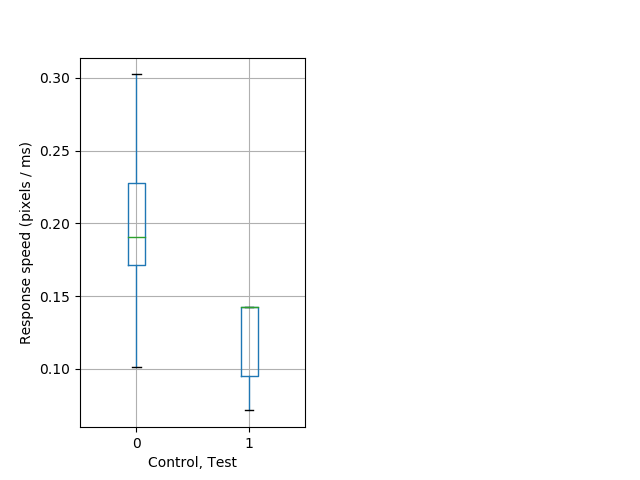
Behavior Data Description 12
Weak tap Graph Pvalue = 0.0135 13 wt vs 15 hom boxgraph_pd__smallmovesribgraph_mean_ribbon_peakspeed_nightprepulseinhibition101.pngBehavior Data Graph 12
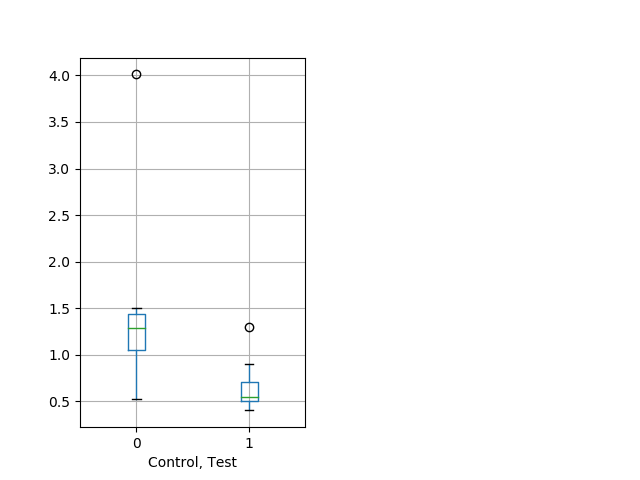
Behavior Data Description 13
NoneBehavior Data Graph 13
NoneBehavior Data Description 14
NoneBehavior Data Graph 14
None